A key region of molecular specificity orchestrates unique ephrin-B1 utilization by Cedar virus
- PMID: 31862858
- PMCID: PMC6925387
- DOI: 10.26508/lsa.201900578
A key region of molecular specificity orchestrates unique ephrin-B1 utilization by Cedar virus
Abstract
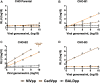
The emergent zoonotic henipaviruses, Hendra, and Nipah are responsible for frequent and fatal disease outbreaks in domestic animals and humans. Specificity of henipavirus attachment glycoproteins (G) for highly species-conserved ephrin ligands underpins their broad host range and is associated with systemic and neurological disease pathologies. Here, we demonstrate that Cedar virus (CedV)-a related henipavirus that is ostensibly nonpathogenic-possesses an idiosyncratic entry receptor repertoire that includes the common henipaviral receptor, ephrin-B2, but, distinct from pathogenic henipaviruses, does not include ephrin-B3. Uniquely among known henipaviruses, CedV can use ephrin-B1 for cellular entry. Structural analyses of CedV-G reveal a key region of molecular specificity that directs ephrin-B1 utilization, while preserving a universal mode of ephrin-B2 recognition. The structural and functional insights presented uncover diversity within the known henipavirus receptor repertoire and suggest that only modest structural changes may be required to modulate receptor specificities within this group of lethal human pathogens.
© 2019 Pryce et al.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures





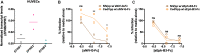



References
-
- WHO (2018) Annual review of diseases prioritized under the research and development blueprint 2018. https://www.who.int/emergencies/diseases/2018prioritization-report.pdf.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
