Precision cancer monitoring using a novel, fully integrated, microfluidic array partitioning digital PCR platform
- PMID: 31862911
- PMCID: PMC6925289
- DOI: 10.1038/s41598-019-55872-7
Precision cancer monitoring using a novel, fully integrated, microfluidic array partitioning digital PCR platform
Abstract
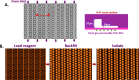
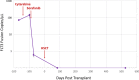
A novel digital PCR (dPCR) platform combining off-the-shelf reagents, a micro-molded plastic microfluidic consumable with a fully integrated single dPCR instrument was developed to address the needs for routine clinical diagnostics. This new platform offers a simplified workflow that enables: rapid time-to-answer; low potential for cross contamination; minimal sample waste; all within a single integrated instrument. Here we showcase the capability of this fully integrated platform to detect and quantify non-small cell lung carcinoma (NSCLC) rare genetic mutants (EGFR T790M) with precision cell-free DNA (cfDNA) standards. Next, we validated the platform with an established chronic myeloid leukemia (CML) fusion gene (BCR-ABL1) assay down to 0.01% mutant allele frequency to highlight the platform's utility for precision cancer monitoring. Thirdly, using a juvenile myelomonocytic leukemia (JMML) patient-specific assay we demonstrate the ability to precisely track an individual cancer patient's response to therapy and show the patient's achievement of complete molecular remission. These three applications highlight the flexibility and utility of this novel fully integrated dPCR platform that has the potential to transform personalized medicine for cancer recurrence monitoring.
Conflict of interest statement
The authors declare no competing interests.
Figures





Similar articles
-
Next-Generation Sequencing-Assisted DNA-Based Digital PCR for a Personalized Approach to the Detection and Quantification of Residual Disease in Chronic Myeloid Leukemia Patients.J Mol Diagn. 2016 Mar;18(2):176-89. doi: 10.1016/j.jmoldx.2015.09.005. Epub 2016 Feb 5. J Mol Diagn. 2016. PMID: 26857065
-
EGFR mutation detection in ctDNA from NSCLC patient plasma: A cross-platform comparison of leading technologies to support the clinical development of AZD9291.Lung Cancer. 2015 Dec;90(3):509-15. doi: 10.1016/j.lungcan.2015.10.004. Epub 2015 Oct 9. Lung Cancer. 2015. PMID: 26494259
-
A Comparison of ddPCR and ARMS for detecting EGFR T790M status in ctDNA from advanced NSCLC patients with acquired EGFR-TKI resistance.Cancer Med. 2017 Jan;6(1):154-162. doi: 10.1002/cam4.978. Epub 2016 Dec 20. Cancer Med. 2017. PMID: 28000387 Free PMC article.
-
Circulating cell-free nucleic acids and platelets as a liquid biopsy in the provision of personalized therapy for lung cancer patients.Lung Cancer. 2017 May;107:100-107. doi: 10.1016/j.lungcan.2016.04.026. Epub 2016 May 4. Lung Cancer. 2017. PMID: 27180141 Review.
-
Emerging digital PCR technology in precision medicine.Biosens Bioelectron. 2022 Sep 1;211:114344. doi: 10.1016/j.bios.2022.114344. Epub 2022 May 14. Biosens Bioelectron. 2022. PMID: 35598553 Review.
Cited by
-
Technical Advances in Circulating Cell-Free DNA Detection and Analysis for Personalized Medicine in Patients' Care.Biomolecules. 2024 Apr 19;14(4):498. doi: 10.3390/biom14040498. Biomolecules. 2024. PMID: 38672514 Free PMC article. Review.
-
CYP2D6 copy number determination using digital PCR.Front Pharmacol. 2024 Aug 14;15:1429286. doi: 10.3389/fphar.2024.1429286. eCollection 2024. Front Pharmacol. 2024. PMID: 39206265 Free PMC article.
-
Advancing Leukemia Management Through Liquid Biopsy: Insights into Biomarkers and Clinical Utility.Cancers (Basel). 2025 Apr 25;17(9):1438. doi: 10.3390/cancers17091438. Cancers (Basel). 2025. PMID: 40361366 Free PMC article. Review.
-
Molybdenum Disulfide as Tunable Electrochemical and Optical Biosensing Platforms for Cancer Biomarker Detection: A Review.Biosensors (Basel). 2023 Aug 25;13(9):848. doi: 10.3390/bios13090848. Biosensors (Basel). 2023. PMID: 37754082 Free PMC article. Review.
-
Robust and rapid partitioning in thermoplastic.Analyst. 2023 Dec 18;149(1):100-107. doi: 10.1039/d3an01869e. Analyst. 2023. PMID: 37982399 Free PMC article.
References
-
- Stetson, D. et al. Orthogonal Comparison of Four Plasma NGS Tests With Tumor Suggests Technical Factors are a Major Source of Assay Discordance. 10.1200/PO.18.00191 (2019). - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

