Divalent cations promote TALE DNA-binding specificity
- PMID: 31863586
- PMCID: PMC7026652
- DOI: 10.1093/nar/gkz1174
Divalent cations promote TALE DNA-binding specificity
Abstract
Recent advances in gene editing have been enabled by programmable nucleases such as transcription activator-like effector nucleases (TALENs) and CRISPR-Cas9. However, several open questions remain regarding the molecular machinery in these systems, including fundamental search and binding behavior as well as role of off-target binding and specificity. In order to achieve efficient and specific cleavage at target sites, a high degree of target site discrimination must be demonstrated for gene editing applications. In this work, we studied the binding affinity and specificity for a series of TALE proteins under a variety of solution conditions using in vitro fluorescence methods and molecular dynamics (MD) simulations. Remarkably, we identified that TALEs demonstrate high sequence specificity only upon addition of small amounts of certain divalent cations (Mg2+, Ca2+). However, under purely monovalent salt conditions (K+, Na+), TALEs bind to specific and non-specific DNA with nearly equal affinity. Divalent cations preferentially bind to DNA over monovalent cations, which attenuates non-specific interactions between TALEs and DNA and further stabilizes specific interactions. Overall, these results uncover new mechanistic insights into the binding action of TALEs and further provide potential avenues for engineering and application of TALE- or TALEN-based systems for genome editing and regulation.
© The Author(s) 2019. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures
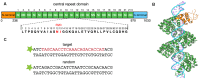
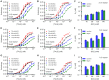
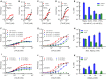
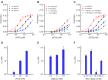
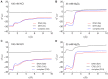
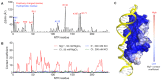
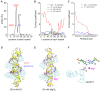
Similar articles
-
A Single-Molecule View of Genome Editing Proteins: Biophysical Mechanisms for TALEs and CRISPR/Cas9.Annu Rev Chem Biomol Eng. 2017 Jun 7;8:577-597. doi: 10.1146/annurev-chembioeng-060816-101603. Epub 2017 May 10. Annu Rev Chem Biomol Eng. 2017. PMID: 28489428 Review.
-
TALEN-Mediated Mutagenesis and Genome Editing.Methods Mol Biol. 2016;1451:17-30. doi: 10.1007/978-1-4939-3771-4_2. Methods Mol Biol. 2016. PMID: 27464798 Free PMC article.
-
Gene Editing With TALEN and CRISPR/Cas in Rice.Prog Mol Biol Transl Sci. 2017;149:81-98. doi: 10.1016/bs.pmbts.2017.04.006. Epub 2017 May 24. Prog Mol Biol Transl Sci. 2017. PMID: 28712502 Review.
-
Non-RVD mutations that enhance the dynamics of the TAL repeat array along the superhelical axis improve TALEN genome editing efficacy.Sci Rep. 2016 Nov 24;6:37887. doi: 10.1038/srep37887. Sci Rep. 2016. PMID: 27883072 Free PMC article.
-
Development of Toolboxes for Precision Genome/Epigenome Editing and Imaging of Epigenetics.Chem Rec. 2018 Dec;18(12):1717-1726. doi: 10.1002/tcr.201800036. Epub 2018 Aug 1. Chem Rec. 2018. PMID: 30066981 Review.
Cited by
-
Role of Divalent Cations in Infections in Host-Pathogen Interaction.Int J Mol Sci. 2024 Sep 10;25(18):9775. doi: 10.3390/ijms25189775. Int J Mol Sci. 2024. PMID: 39337264 Free PMC article. Review.
-
Epigenome engineering: new technologies for precision medicine.Nucleic Acids Res. 2020 Dec 16;48(22):12453-12482. doi: 10.1093/nar/gkaa1000. Nucleic Acids Res. 2020. PMID: 33196851 Free PMC article.
-
Epigenetic Regulation of β-Globin Genes and the Potential to Treat Hemoglobinopathies through Epigenome Editing.Genes (Basel). 2023 Feb 25;14(3):577. doi: 10.3390/genes14030577. Genes (Basel). 2023. PMID: 36980849 Free PMC article. Review.
-
Designing Epigenome Editors: Considerations of Biochemical and Locus Specificities.Methods Mol Biol. 2024;2842:23-55. doi: 10.1007/978-1-0716-4051-7_2. Methods Mol Biol. 2024. PMID: 39012589
-
Tuning up Transcription Factors for Therapy.Molecules. 2020 Apr 20;25(8):1902. doi: 10.3390/molecules25081902. Molecules. 2020. PMID: 32326099 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

