Tracing metabolic flux through time and space with isotope labeling experiments
- PMID: 31864070
- PMCID: PMC7302994
- DOI: 10.1016/j.copbio.2019.11.003
Tracing metabolic flux through time and space with isotope labeling experiments
Abstract
Metabolism is dynamic and must function in context-specific ways to adjust to changes in the surrounding cellular and ecological environment. When isotopic tracers are used, metabolite flow (i.e. metabolic flux) can be quantified through biochemical networks to assess metabolic pathway operation. The cellular activities considered across multiple tissues and organs result in the observed phenotype and can be analyzed to discover emergent, whole-system properties of biology and elucidate misconceptions about network operation. However, temporal and spatial challenges remain significant hurdles and require novel approaches and creative solutions. We survey current investigations in higher plant and animal systems focused on dynamic isotope labeling experiments, spatially resolved measurement strategies, and observations from re-analysis of our own studies that suggest prospects for future work. Related discoveries will be necessary to push the frontier of our understanding of metabolism to suggest novel solutions to cure disease and feed a growing future world population.
Published by Elsevier Ltd.
Conflict of interest statement
Declaration of interests
☒ The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures


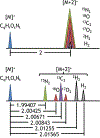
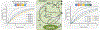
References
-
- Long SP, Marshall-Colon A, Zhu XG: Meeting the global food demand of the future by engineering crop photosynthesis and yield potential. Cell 2015, 161:56–66. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

