Restricted Clonality and Limited Germinal Center Reentry Characterize Memory B Cell Reactivation by Boosting
- PMID: 31866068
- PMCID: PMC6958527
- DOI: 10.1016/j.cell.2019.11.032
Restricted Clonality and Limited Germinal Center Reentry Characterize Memory B Cell Reactivation by Boosting
Abstract
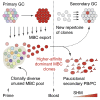
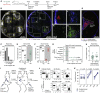
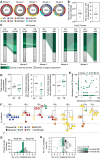
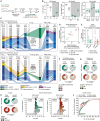
Repeated exposure to pathogens or their antigens triggers anamnestic antibody responses that are higher in magnitude and affinity than the primary response. These involve reengagement of memory B cell (MBC) clones, the diversity and specificity of which determine the breadth and effectiveness of the ensuing antibody response. Using prime-boost models in mice, we find that secondary responses are characterized by a clonality bottleneck that restricts the engagement of the large diversity of MBC clones generated by priming. Rediversification of mutated MBCs is infrequent within secondary germinal centers (GCs), which instead consist predominantly of B cells without prior GC experience or detectable clonal expansion. Few MBC clones, generally derived from higher-affinity germline precursors, account for the majority of secondary antibody responses, while most primary-derived clonal diversity is not reengaged detectably by boosting. Understanding how to counter this bottleneck may improve our ability to elicit antibodies to non-immunodominant epitopes by vaccination.
Keywords: affinity maturation; antibody response; clonal dynamics; germinal center memory B cell; immunological memory; influenza A virus; memory reactivation.
Copyright © 2019 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
The authors declare no competing interests.
Figures




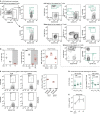


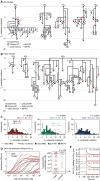
Comment in
-
It's Hard to Teach an Old B Cell New Tricks.Cell. 2020 Jan 9;180(1):18-20. doi: 10.1016/j.cell.2019.12.019. Cell. 2020. PMID: 31951517
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

