Population structure, connectivity, and demographic history of an apex marine predator, the bull shark Carcharhinus leucas
- PMID: 31871624
- PMCID: PMC6912899
- DOI: 10.1002/ece3.5597
Population structure, connectivity, and demographic history of an apex marine predator, the bull shark Carcharhinus leucas
Abstract
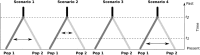
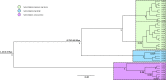
Knowledge of population structure, connectivity, and effective population size remains limited for many marine apex predators, including the bull shark Carcharhinus leucas. This large-bodied coastal shark is distributed worldwide in warm temperate and tropical waters, and uses estuaries and rivers as nurseries. As an apex predator, the bull shark likely plays a vital ecological role within marine food webs, but is at risk due to inshore habitat degradation and various fishing pressures. We investigated the bull shark's global population structure and demographic history by analyzing the genetic diversity of 370 individuals from 11 different locations using 25 microsatellite loci and three mitochondrial genes (CR, nd4, and cytb). Both types of markers revealed clustering between sharks from the Western Atlantic and those from the Western Pacific and the Western Indian Ocean, with no contemporary gene flow. Microsatellite data suggested low differentiation between the Western Indian Ocean and the Western Pacific, but substantial differentiation was found using mitochondrial DNA. Integrating information from both types of markers and using Bayesian computation with a random forest procedure (ABC-RF), this discordance was found to be due to a complete lack of contemporary gene flow. High genetic connectivity was found both within the Western Indian Ocean and within the Western Pacific. In conclusion, these results suggest important structuring of bull shark populations globally with important gene flow occurring along coastlines, highlighting the need for management and conservation plans on regional scales rather than oceanic basin scale.
Keywords: ABC‐RF; microsatellite DNA; mitochondrial DNA; mito‐nuclear discordance; population genetics.
© 2019 The Authors. Ecology and Evolution published by John Wiley & Sons Ltd.
Conflict of interest statement
None declared.
Figures




References
-
- Aguilar, C. , Gonzalez‐Sanson, G. , Hueter, R. , Rojas, E. , Cabrera, Y. , Briones, A. , … Baker, P. (2014). Shark catches in the northwest region of Cuba. Latin American Journal of Aquatic Research, 42(3), 477–487.
-
- Almeida, O. T. , McGrath, D. G. , & Ruffino, M. L. (2001). The commercial fisheries of the lower Amazon: An economic analysis. Fisheries Management and Ecology, 8(3), 253–269. 10.1046/j.1365-2400.2001.00234.x - DOI
-
- Arevalo, E. , Davis, S. K. , & Sites, J. W. (1994). Mitochondrial‐DNA sequence divergence and phylogenetic‐relationships among 8 chromosome races of the Sceloporus‐Grammicus Complex (Phrynosomatidae) in Central Mexico. Systematic Biology, 43(3), 387–418. 10.2307/2413675 - DOI
-
- Bailleul, D. , Mackenzie, A. , Sacchi, O. , Poisson, F. , Bierne, N. , & Arnaud‐Haond, S. (2018). Large‐scale genetic panmixia in the blue shark (Prionace glauca): A single worldwide population, or a genetic lag‐time effect of the “grey zone” of differentiation? Evolutionary Applications, 11(5), 614–630. 10.1111/eva.12591 - DOI - PMC - PubMed
-
- Beaumont, M. A. (2010). Approximate Bayesian Computation in evolution and ecology. Annual Review of Ecology, Evolution, and Systematics, 41(1), 379–406. 10.1146/annurev-ecolsys-102209-144621 - DOI
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

