On ATG4B as Drug Target for Treatment of Solid Tumours-The Knowns and the Unknowns
- PMID: 31878323
- PMCID: PMC7016753
- DOI: 10.3390/cells9010053
On ATG4B as Drug Target for Treatment of Solid Tumours-The Knowns and the Unknowns
Abstract
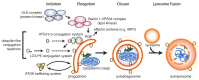
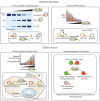
Autophagy is an evolutionary conserved stress survival pathway that has been shown to play an important role in the initiation, progression, and metastasis of multiple cancers; however, little progress has been made to date in translation of basic research to clinical application. This is partially due to an incomplete understanding of the role of autophagy in the different stages of cancer, and also to an incomplete assessment of potential drug targets in the autophagy pathway. While drug discovery efforts are on-going to target enzymes involved in the initiation phase of the autophagosome, e.g., unc51-like autophagy activating kinase (ULK)1/2, vacuolar protein sorting 34 (Vps34), and autophagy-related (ATG)7, we propose that the cysteine protease ATG4B is a bona fide drug target for the development of anti-cancer treatments. In this review, we highlight some of the recent advances in our understanding of the role of ATG4B in autophagy and its relevance to cancer, and perform a critical evaluation of ATG4B as a druggable cancer target.
Keywords: ATG4; autophagy; biomarker; drug screening; pancreatic ductal adenocarcinoma (PDAC); screening assay; small molecule compound.
Conflict of interest statement
The authors declare no conflicts of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Figures

References
-
- Amaravadi R.K., Lippincott-Schwartz J., Yin X.M., Weiss W.A., Takebe N., Timmer W., DiPaola R.S., Lotze M.T., White E. Principles and current strategies for targeting autophagy for cancer treatment. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2011;17:654–666. doi: 10.1158/1078-0432.CCR-10-2634. - DOI - PMC - PubMed
-
- Amaravadi R.K., Kimmelman A.C., Debnath J. Targeting Autophagy in Cancer: Recent Advances and Future Directions. Cancer Discov. 2019;9:1167–1181. doi: 10.1158/2159-8290.CD-19-0292. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

