Multivalent interaction of ESCO2 with the replication machinery is required for sister chromatid cohesion in vertebrates
- PMID: 31879348
- PMCID: PMC6969535
- DOI: 10.1073/pnas.1911936117
Multivalent interaction of ESCO2 with the replication machinery is required for sister chromatid cohesion in vertebrates
Abstract
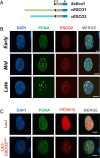
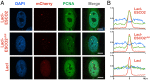
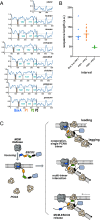
The tethering together of sister chromatids by the cohesin complex ensures their accurate alignment and segregation during cell division. In vertebrates, sister chromatid cohesion requires the activity of the ESCO2 acetyltransferase, which modifies the Smc3 subunit of cohesin. It was shown recently that ESCO2 promotes cohesion through interaction with the MCM replicative helicase. However, ESCO2 does not significantly colocalize with the MCM complex, suggesting there are additional interactions important for ESCO2 function. Here we show that ESCO2 is recruited to replication factories, sites of DNA replication, through interaction with PCNA. We show that ESCO2 contains multiple PCNA-interaction motifs in its N terminus, each of which is essential to its ability to establish cohesion. We propose that multiple PCNA-interaction motifs embedded in a largely flexible and disordered region of the protein underlie the unique ability of ESCO2 to establish cohesion between sister chromatids precisely as they are born during DNA replication.
Keywords: DNA replication; chromosome biology; cohesin; sister chromatid cohesion.
Conflict of interest statement
The authors declare no competing interest.
Figures



References
-
- Gerlich D., Koch B., Dupeux F., Peters J.-M., Ellenberg J., Live-cell imaging reveals a stable cohesin-chromatin interaction after but not before DNA replication. Curr. Biol. 16, 1571–1578 (2006). - PubMed
-
- Zhang J., et al. , Acetylation of Smc3 by Eco1 is required for S phase sister chromatid cohesion in both human and yeast. Mol. Cell 31, 143–151 (2008). - PubMed
-
- Sutani T., Kawaguchi T., Kanno R., Itoh T., Shirahige K., Budding yeast Wpl1(Rad61)-Pds5 complex counteracts sister chromatid cohesion-establishing reaction. Curr. Biol. 19, 492–497 (2009). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

