Determinants governing T cell receptor α/β-chain pairing in repertoire formation of identical twins
- PMID: 31879353
- PMCID: PMC6955297
- DOI: 10.1073/pnas.1915008117
Determinants governing T cell receptor α/β-chain pairing in repertoire formation of identical twins
Abstract
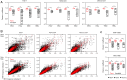
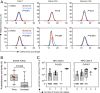
The T cell repertoire in each individual includes T cell receptors (TCRs) of enormous sequence diversity through the pairing of diverse TCR α- and β-chains, each generated by somatic recombination of paralogous gene segments. Whether the TCR repertoire contributes to susceptibility to infectious or autoimmune diseases in concert with disease-associated major histocompatibility complex (MHC) polymorphisms is unknown. Due to a lack in high-throughput technologies to sequence TCR α-β pairs, current studies on whether the TCR repertoire is shaped by host genetics have so far relied only on single-chain analysis. Using a high-throughput single T cell sequencing technology, we obtained the largest paired TCRαβ dataset so far, comprising 965,523 clonotypes from 15 healthy individuals including 6 monozygotic twin pairs. Public TCR α- and, to a lesser extent, TCR β-chain sequences were common in all individuals. In contrast, sharing of entirely identical TCRαβ amino acid sequences was very infrequent in unrelated individuals, but highly increased in twins, in particular in CD4 memory T cells. Based on nucleotide sequence identity, a subset of these shared clonotypes appeared to be the progeny of T cells that had been generated during fetal development and had persisted for more than 50 y. Additional shared TCRαβ in twins were encoded by different nucleotide sequences, implying that genetic determinants impose structural constraints on thymic selection that favor the selection of TCR α-β pairs with entire sequence identities.
Keywords: T cell repertoire; major histocompatibility complex; monozygotic twins; single-cell sequencing.
Conflict of interest statement
The authors declare no competing interest.
Figures


References
-
- Yoshikai Y., et al. , Organization and sequences of the variable, joining and constant region genes of the human T-cell receptor alpha-chain. Nature 316, 837–840 (1985). - PubMed
-
- Arstila T. P., et al. , A direct estimate of the human alphabeta T cell receptor diversity. Science 286, 958–961 (1999). - PubMed
-
- Arstila T. P., et al. , Diversity of human alpha beta T cell receptors. Science 288, 1135 (2000). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

