Hierarchical Hybrid Enrichment: Multitiered Genomic Data Collection Across Evolutionary Scales, With Application to Chorus Frogs (Pseudacris)
- PMID: 31886503
- PMCID: PMC7302053
- DOI: 10.1093/sysbio/syz074
Hierarchical Hybrid Enrichment: Multitiered Genomic Data Collection Across Evolutionary Scales, With Application to Chorus Frogs (Pseudacris)
Abstract
Determining the optimal targets of genomic subsampling for phylogenomics, phylogeography, and population genomics remains a challenge for evolutionary biologists. Of the available methods for subsampling the genome, hybrid enrichment (sequence capture) has become one of the primary means of data collection for systematics, due to the flexibility and cost efficiency of this approach. Despite the utility of this method, information is lacking as to what genomic targets are most appropriate for addressing questions at different evolutionary scales. In this study, first, we compare the benefits of target loci developed for deep- and shallow scales by comparing these loci at each of three taxonomic levels: within a genus (phylogenetics), within a species (phylogeography), and within a hybrid zone (population genomics). Specifically, we target evolutionarily conserved loci that are appropriate for deeper phylogenetic scales and more rapidly evolving loci that are informative for phylogeographic and population genomic scales. Second, we assess the efficacy of targeting multiple-locus sets for different taxonomic levels in the same hybrid enrichment reaction, an approach we term hierarchical hybrid enrichment. Third, we apply this approach to the North American chorus frog genus Pseudacris to answer key evolutionary questions across taxonomic and temporal scales. We demonstrate that in this system the type of genomic target that produces the most resolved gene trees differs depending on the taxonomic level, although the potential for error is substantially lower for the deep-scale loci at all levels. We successfully recover data for the two different locus sets with high efficiency. Using hierarchical data targeting deep and shallow levels: we 1) resolve the phylogeny of the genus Pseudacris and introduce a novel visual and hypothesis testing method that uses nodal heat maps to examine the robustness of branch support values to the removal of sites and loci; 2) estimate the phylogeographic history of Pseudacris feriarum, which reveals up to five independent invasions leading to sympatry with congener Pseudacris nigrita to form replicated reinforcement contact zones with ongoing gene flow into sympatry; and 3) quantify with high confidence the frequency of hybridization in one of these zones between P. feriarum and P. nigrita, which is lower than microsatellite-based estimates. We find that the hierarchical hybrid enrichment approach offers an efficient, multitiered data collection method for simultaneously addressing questions spanning multiple evolutionary scales. [Anchored hybrid enrichment; heat map; hybridization; phylogenetics; phylogeography; population genomics; reinforcement; reproductive character displacement.].
© The Author(s) 2019. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For permissions, please email: journals.permissions@oup.com.
Figures




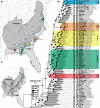
References
-
- Ané C., Larget B., Baum D.A., Smith S.D., Rokas A.. 2007. Bayesian estimation of concordance among gene trees. Mol. Biol. Evol. 24:412–426. - PubMed
-
- Arnold B., Corbett-Detig R.B., Hartl D., Bomblies K.. 2013. RADseq underestimates diversity and introduces genealogical biases due to nonrandom haplotype sampling. Mol. Ecol. 22:3179–3190. - PubMed

