Interpreting pathways to discover cancer driver genes with Moonlight
- PMID: 31900418
- PMCID: PMC6941958
- DOI: 10.1038/s41467-019-13803-0
Interpreting pathways to discover cancer driver genes with Moonlight
Abstract
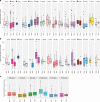
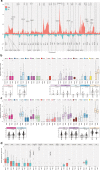
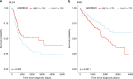
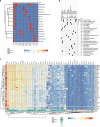
Cancer driver gene alterations influence cancer development, occurring in oncogenes, tumor suppressors, and dual role genes. Discovering dual role cancer genes is difficult because of their elusive context-dependent behavior. We define oncogenic mediators as genes controlling biological processes. With them, we classify cancer driver genes, unveiling their roles in cancer mechanisms. To this end, we present Moonlight, a tool that incorporates multiple -omics data to identify critical cancer driver genes. With Moonlight, we analyze 8000+ tumor samples from 18 cancer types, discovering 3310 oncogenic mediators, 151 having dual roles. By incorporating additional data (amplification, mutation, DNA methylation, chromatin accessibility), we reveal 1000+ cancer driver genes, corroborating known molecular mechanisms. Additionally, we confirm critical cancer driver genes by analysing cell-line datasets. We discover inactivation of tumor suppressors in intron regions and that tissue type and subtype indicate dual role status. These findings help explain tumor heterogeneity and could guide therapeutic decisions.
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Revealing cancer driver genes through integrative transcriptomic and epigenomic analyses with Moonlight.PLoS Comput Biol. 2025 Apr 21;21(4):e1012999. doi: 10.1371/journal.pcbi.1012999. eCollection 2025 Apr. PLoS Comput Biol. 2025. PMID: 40258059 Free PMC article.
-
Identifying cancer type specific oncogenes and tumor suppressors using limited size data.J Bioinform Comput Biol. 2016 Dec;14(6):1650031. doi: 10.1142/S0219720016500311. Epub 2016 Aug 31. J Bioinform Comput Biol. 2016. PMID: 27712196
-
SomInaClust: detection of cancer genes based on somatic mutation patterns of inactivation and clustering.BMC Bioinformatics. 2015 Apr 23;16:125. doi: 10.1186/s12859-015-0555-7. BMC Bioinformatics. 2015. PMID: 25903787 Free PMC article.
-
The cancer driver genes IDH1/2, JARID1C/ KDM5C, and UTX/ KDM6A: crosstalk between histone demethylation and hypoxic reprogramming in cancer metabolism.Exp Mol Med. 2019 Jun 20;51(6):1-17. doi: 10.1038/s12276-019-0230-6. Exp Mol Med. 2019. PMID: 31221981 Free PMC article. Review.
-
The INPP4B paradox: Like PTEN, but different.Adv Biol Regul. 2021 Dec;82:100817. doi: 10.1016/j.jbior.2021.100817. Epub 2021 Jun 16. Adv Biol Regul. 2021. PMID: 34216856 Review.
Cited by
-
EPIMUTESTR: a nearest neighbor machine learning approach to predict cancer driver genes from the evolutionary action of coding variants.Nucleic Acids Res. 2022 Jul 8;50(12):e70. doi: 10.1093/nar/gkac215. Nucleic Acids Res. 2022. PMID: 35412634 Free PMC article.
-
Comprehensive evaluation of computational methods for predicting cancer driver genes.Brief Bioinform. 2022 Mar 10;23(2):bbab548. doi: 10.1093/bib/bbab548. Brief Bioinform. 2022. PMID: 35037014 Free PMC article.
-
A molecular taxonomy of tumors independent of tissue-of-origin.iScience. 2021 Sep 3;24(10):103084. doi: 10.1016/j.isci.2021.103084. eCollection 2021 Oct 22. iScience. 2021. PMID: 34611608 Free PMC article.
-
Integrated Proteogenomic Characterization across Major Histological Types of Pediatric Brain Cancer.Cell. 2020 Dec 23;183(7):1962-1985.e31. doi: 10.1016/j.cell.2020.10.044. Epub 2020 Nov 25. Cell. 2020. PMID: 33242424 Free PMC article.
-
Proteogenomic insights into the biology and treatment of HPV-negative head and neck squamous cell carcinoma.Cancer Cell. 2021 Mar 8;39(3):361-379.e16. doi: 10.1016/j.ccell.2020.12.007. Epub 2021 Jan 7. Cancer Cell. 2021. PMID: 33417831 Free PMC article.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

