Disrupted ER membrane protein complex-mediated topogenesis drives congenital neural crest defects
- PMID: 31904590
- PMCID: PMC6994125
- DOI: 10.1172/JCI129308
Disrupted ER membrane protein complex-mediated topogenesis drives congenital neural crest defects
Abstract
Multipass membrane proteins have a myriad of functions, including transduction of cell-cell signals, ion transport, and photoreception. Insertion of these proteins into the membrane depends on the endoplasmic reticulum (ER) membrane protein complex (EMC). Recently, birth defects have been observed in patients with variants in the gene encoding a member of this complex, EMC1. Patient phenotypes include congenital heart disease, craniofacial malformations, and neurodevelopmental disease. However, a molecular connection between EMC1 and these birth defects is lacking. Using Xenopus, we identified defects in neural crest cells (NCCs) upon emc1 depletion. We then used unbiased proteomics and discovered a critical role for emc1 in WNT signaling. Consistent with this, readouts of WNT signaling and Frizzled (Fzd) levels were reduced in emc1-depleted embryos, while NCC defects could be rescued with β-catenin. Interestingly, other transmembrane proteins were mislocalized upon emc1 depletion, providing insight into additional patient phenotypes. To translate our findings back to humans, we found that EMC1 was necessary for human NCC development in vitro. Finally, we tested patient variants in our Xenopus model and found the majority to be loss-of-function alleles. Our findings define molecular mechanisms whereby EMC1 dysfunction causes disease phenotypes through dysfunctional multipass membrane protein topogenesis.
Keywords: Cardiovascular disease; Development; Embryonic development; Genetics; Monogenic diseases.
Conflict of interest statement
Figures

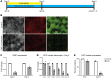

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

