Long-term stability of cortical population dynamics underlying consistent behavior
- PMID: 31907438
- PMCID: PMC7007364
- DOI: 10.1038/s41593-019-0555-4
Long-term stability of cortical population dynamics underlying consistent behavior
Abstract
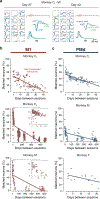
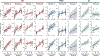
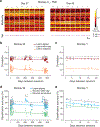
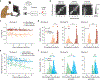
Animals readily execute learned behaviors in a consistent manner over long periods of time, and yet no equally stable neural correlate has been demonstrated. How does the cortex achieve this stable control? Using the sensorimotor system as a model of cortical processing, we investigated the hypothesis that the dynamics of neural latent activity, which captures the dominant co-variation patterns within the neural population, must be preserved across time. We recorded from populations of neurons in premotor, primary motor and somatosensory cortices as monkeys performed a reaching task, for up to 2 years. Intriguingly, despite a steady turnover in the recorded neurons, the low-dimensional latent dynamics remained stable. The stability allowed reliable decoding of behavioral features for the entire timespan, while fixed decoders based directly on the recorded neural activity degraded substantially. We posit that stable latent cortical dynamics within the manifold are the fundamental building blocks underlying consistent behavioral execution.
Conflict of interest statement
COMPETING INTERESTS STATEMENT
The authors declare no competing interests.
Figures



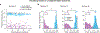




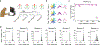





References
-
- Stopfer M, Jayaraman V & Laurent G. Intensity versus Identity Coding in an Olfactory System. Neuron 39, 991–1004 (2003). - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

