High-resolution mapping of reciprocal translocation breakpoints using long-read sequencing
- PMID: 31908979
- PMCID: PMC6939040
- DOI: 10.1016/j.mex.2019.10.028
High-resolution mapping of reciprocal translocation breakpoints using long-read sequencing
Abstract
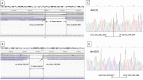
Long-read nanopore sequencing enables direct high-resolution breakpoint mapping on balanced carriers of reciprocal translocation. The mean sequencing depth on the translocated chromosomes to achieve accurate mapping of breakpoints ranged from 2.5-fold to 6.2-fold. To speed up determination of the breakpoints from long-read sequencing data, alignment reads on the translocated chromosomes were extracted before piped into NanoSV. Checking the position of breakpoints on Interactive Genomics Viewer (IGV) was crucial to successful design of breakpoint PCR primers, especially when large deletion was involved at the breakpoints. •Long-read sequencing enables accurate breakpoint mapping with base-pair resolution•Splitting bam files by translocated chromosomes drastically speeded up the breakpoint determination•IGV helps to identify the breakpoint positions and facilitate the design of breakpoint PCR primers.
Keywords: Breakpoint; Nanopore sequencing; PGT-SR; Translocation breakpoint mapping using nanopore sequencing.
© 2019 The Author(s).
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures


Similar articles
-
Distinguishing between carrier and noncarrier embryos with the use of long-read sequencing in preimplantation genetic testing for reciprocal translocations.Genomics. 2020 Jan;112(1):494-500. doi: 10.1016/j.ygeno.2019.04.001. Epub 2019 Apr 1. Genomics. 2020. PMID: 30946890
-
Analysis of Preimplantation and Clinical Outcomes of Two Cases by Oxford Nanopore Sequencing.Reprod Sci. 2024 Jul;31(7):2123-2134. doi: 10.1007/s43032-024-01470-6. Epub 2024 Feb 12. Reprod Sci. 2024. PMID: 38347380
-
Location of Balanced Chromosome-Translocation Breakpoints by Long-Read Sequencing on the Oxford Nanopore Platform.Front Genet. 2020 Jan 14;10:1313. doi: 10.3389/fgene.2019.01313. eCollection 2019. Front Genet. 2020. PMID: 32010185 Free PMC article.
-
Translocation breakpoints of chromosome 3 in male carriers: a report of twelve cases and a review of the literature.Turk J Med Sci. 2018 Feb 23;48(1):150-156. doi: 10.3906/sag-1704-1. Turk J Med Sci. 2018. PMID: 29479975 Review.
-
Historical and Clinical Perspectives on Chromosomal Translocations.Adv Exp Med Biol. 2018;1044:1-14. doi: 10.1007/978-981-13-0593-1_1. Adv Exp Med Biol. 2018. PMID: 29956287 Review.
Cited by
-
Mate-pair sequencing assisted prenatal counseling for a rare complex chromosomal rearrangement carrier.Hum Mol Genet. 2025 May 6;34(10):864-869. doi: 10.1093/hmg/ddaf012. Hum Mol Genet. 2025. PMID: 40044118 Free PMC article.
-
Nanopore sequencing with T2T-CHM13 for accurate detection and preventing the transmission of structural rearrangements in highly repetitive heterochromatin regions in human embryos.Clin Transl Med. 2024 Mar;14(3):e1612. doi: 10.1002/ctm2.1612. Clin Transl Med. 2024. PMID: 38445430 Free PMC article.
-
Gene sequencing and result analysis of balanced translocation carriers by third-generation gene sequencing technology.Sci Rep. 2023 Apr 28;13(1):7004. doi: 10.1038/s41598-022-20356-8. Sci Rep. 2023. PMID: 37117255 Free PMC article.
References
-
- Chow J.F.C., Cheng H.H.Y., Lau E.Y.L., Yeung W.S.B., Ng E.H.Y. Distinguishing between carrier and noncarrier embryos with the use of long-read sequencing in preimplantation genetic testing for reciprocal translocations. Genomics. 2019 - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

