Involvement of Rev1 in alkylating agent-induced loss of heterozygosity in Oryzias latipes
- PMID: 31917895
- PMCID: PMC7079036
- DOI: 10.1111/gtc.12746
Involvement of Rev1 in alkylating agent-induced loss of heterozygosity in Oryzias latipes
Abstract
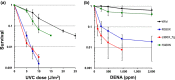
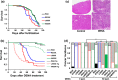
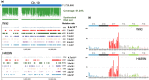
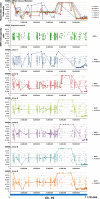
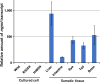
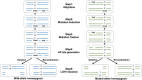
Translesion synthesis (TLS) polymerases mediate DNA damage bypass during replication. The TLS polymerase Rev1 has two important functions in the TLS pathway, including dCMP transferase activity and acting as a scaffolding protein for other TLS polymerases at the C-terminus. Because of the former activity, Rev1 bypasses apurinic/apyrimidinic sites by incorporating dCMP, whereas the latter activity mediates assembly of multipolymerase complexes at the DNA lesions. We generated rev1 mutants lacking each of these two activities in Oryzias latipes (medaka) fish and analyzed cytotoxicity and mutagenicity in response to the alkylating agent diethylnitrosamine (DENA). Mutant lacking the C-terminus was highly sensitive to DENA cytotoxicity, whereas mutant with reduced dCMP transferase activity was slightly sensitive to DENA cytotoxicity, but exhibited a higher tumorigenic rate than wild-type fish. There was no significant difference in the frequency of DENA-induced mutations between mutant with reduced dCMP transferase activity and wild-type cultured cell. However, loss of heterozygosity (LOH) occurred frequently in cells with reduced dCMP transferase activity. LOH is a common genetic event in many cancer types and plays an important role on carcinogenesis. To our knowledge, this is the first report to identify the involvement of the catalytic activity of Rev1 in suppression of LOH.
Keywords: DNA damage; DNA polymerase; alkylating agent; chemical mutagenesis; loss of heterozygosity; mutagenesis; translesion synthesis.
© 2020 The Authors. Genes to Cells published by Molecular Biology Society of Japan and John Wiley & Sons Australia, Ltd.
Conflict of interest statement
The authors declare no competing financial interests.
Figures


Similar articles
-
Rev1 overexpression accelerates N-methyl-N-nitrosourea (MNU)-induced thymic lymphoma by increasing mutagenesis.Cancer Sci. 2024 Jun;115(6):1808-1819. doi: 10.1111/cas.16159. Epub 2024 Apr 4. Cancer Sci. 2024. PMID: 38572512 Free PMC article.
-
The catalytic function of the Rev1 dCMP transferase is required in a lesion-specific manner for translesion synthesis and base damage-induced mutagenesis.Nucleic Acids Res. 2010 Aug;38(15):5036-46. doi: 10.1093/nar/gkq225. Epub 2010 Apr 12. Nucleic Acids Res. 2010. PMID: 20388628 Free PMC article.
-
A non-catalytic function of Rev1 in translesion DNA synthesis and mutagenesis is mediated by its stable interaction with Rad5.DNA Repair (Amst). 2013 Jan 1;12(1):27-37. doi: 10.1016/j.dnarep.2012.10.003. Epub 2012 Nov 9. DNA Repair (Amst). 2013. PMID: 23142547
-
Roles of mutagenic translesion synthesis in mammalian genome stability, health and disease.DNA Repair (Amst). 2015 May;29:56-64. doi: 10.1016/j.dnarep.2015.01.001. Epub 2015 Jan 21. DNA Repair (Amst). 2015. PMID: 25655219 Review.
-
Translesion synthesis by the UmuC family of DNA polymerases.Mutat Res. 2001 Jul 12;486(2):59-70. doi: 10.1016/s0921-8777(01)00089-1. Mutat Res. 2001. PMID: 11425512 Review.
Cited by
-
Rev1 overexpression accelerates N-methyl-N-nitrosourea (MNU)-induced thymic lymphoma by increasing mutagenesis.Cancer Sci. 2024 Jun;115(6):1808-1819. doi: 10.1111/cas.16159. Epub 2024 Apr 4. Cancer Sci. 2024. PMID: 38572512 Free PMC article.
-
Targeting translesion synthesis (TLS) to expose replication gaps, a unique cancer vulnerability.Expert Opin Ther Targets. 2021 Jan;25(1):27-36. doi: 10.1080/14728222.2021.1864321. Epub 2021 Jan 8. Expert Opin Ther Targets. 2021. PMID: 33416413 Free PMC article. Review.
References
-
- Auerbach, P. , Bennett, R. A. , Bailey, E. A. , Krokan, H. E. , & Demple, B. (2005). Mutagenic specificity of endogenously generated abasic sites in Saccharomyces cerevisiae chromosomal DNA. Proceedings of the National Academy of Sciences of the United States of America, 102, 17711–17716. 10.1073/pnas.0504643102 - DOI - PMC - PubMed
-
- Brown‐Peterson, N. J. , Krol, R. M. , Zhu, Y. L. , & Hawkins, W. E. (1999). N‐nitrosodiethylamine initiation of carcinogenesis in Japanese medaka (Oryzias latipes): Hepatocellular proliferation, toxicity, and neoplastic lesions resulting from short term, low level exposure. Toxicological Sciences, 50, 186–194. 10.1093/toxsci/50.2.186 - DOI - PubMed
-
- Bubenshchikova, E. , Ju, B. , Pristyazhnyuk, I. , Niwa, K. , Kaftanovskaya, E. , Kinoshita, M. , … Wakamatsu, Y. (2005). Generation of fertile and diploid fish, medaka (Oryzias latipes), from nuclear transplantation of blastula and four‐somite‐stage embryonic cells into nonenucleated unfertilized eggs. Cloning and Stem Cells, 7, 255–264. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

