NXF1 and CRM1 nuclear export pathways orchestrate nuclear export, translation and packaging of murine leukaemia retrovirus unspliced RNA
- PMID: 31918596
- PMCID: PMC7237160
- DOI: 10.1080/15476286.2020.1713539
NXF1 and CRM1 nuclear export pathways orchestrate nuclear export, translation and packaging of murine leukaemia retrovirus unspliced RNA
Abstract
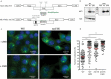
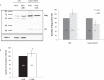
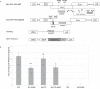
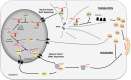
Cellular mRNAs are exported from the nucleus as fully spliced RNAs. Proofreading mechanisms eliminate unprocessed and irregular pre-mRNAs to control the quality of gene expression. Retroviruses need to export partially spliced and unspliced full-length RNAs to the cytoplasm where they serve as templates for protein synthesis and/or as encapsidated RNA in progeny viruses. Genetically complex retroviruses such as HIV-1 use Rev-equivalent proteins to export intron-retaining RNA from the nucleus using the cellular CRM1-driven nuclear export machinery. By contrast, genetically simpler retroviruses such as murine leukaemia virus (MLV) recruit the NXF1 RNA export machinery. In this study, we reveal for the first time that MLV hijacks both NXF1 and CRM1-dependent pathways to achieve optimal replication capacity. The CRM1-pathway marks the MLV full-length RNA (FL RNA) for packaging, while NXF1-driven nuclear export is coupled to translation. Thus, the cytoplasmic function of the viral RNA is determined early in the nucleus. Depending on the nature of ribonucleoprotein complex formed on FL RNA cargo in the nucleus, the FL RNA will be addressed to the translation machinery sites or to the virus-assembly sites at the plasma membrane.
Keywords: CRM1; MLV; NXF1/Tap; RNA; export; packaging; retrovirus; trafficking.
Figures

References
-
- Kanopka A, Muhlemann O, Akusjarvi G.. Inhibition by SR proteins splicing of a regulated adenovirus pre-mRNA. Nature. 1996;381:535–538. - PubMed
-
- Jang SK, Wimmer E. Cap-independent translation of encephalomyocarditis virus RNA: structural elements of the internal ribosomal entry site and involvement of a cellular 57-kD RNA-binding protein. Genes Dev. 1990;4:1560–1572. - PubMed
-
- Fornerod M, Ohno M, Yoshida M, et al. CRM1 is an export receptor for leucine-rich nuclear export signals. Cell. 1997;90:1051–1060. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
