A fluorescence-activatable reporter of flavivirus NS2B-NS3 protease activity enables live imaging of infection in single cells and viral plaques
- PMID: 31919100
- PMCID: PMC7039541
- DOI: 10.1074/jbc.RA119.011319
A fluorescence-activatable reporter of flavivirus NS2B-NS3 protease activity enables live imaging of infection in single cells and viral plaques
Abstract
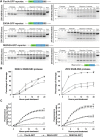
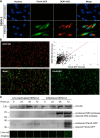
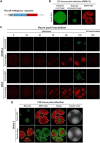
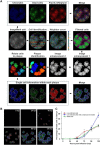
The genus Flavivirus in the family Flaviviridae comprises many medically important viruses, such as dengue virus (DENV), Zika virus (ZIKV), and yellow fever virus. The quest for therapeutic targets to combat flavivirus infections requires a better understanding of the kinetics of virus-host interactions during infections with native viral strains. However, this is precluded by limitations of current cell-based systems for monitoring flavivirus infection in living cells. In the present study, we report the construction of fluorescence-activatable sensors to detect the activities of flavivirus NS2B-NS3 serine proteases in living cells. The system consists of GFP-based reporters that become fluorescent upon cleavage by recombinant DENV-2/ZIKV proteases in vitro A version of this sensor containing the flavivirus internal NS3 cleavage site linker reported the highest fluorescence activation in stably transduced mammalian cells upon DENV-2/ZIKV infection. Moreover, the onset of fluorescence correlated with viral protease activity. A far-red version of this flavivirus sensor had the best signal-to-noise ratio in a fluorescent Dulbecco's plaque assay, leading to the construction of a multireporter platform combining the flavivirus sensor with reporter dyes for detection of chromatin condensation and cell death, enabling studies of viral plaque formation with single-cell resolution. Finally, the application of this platform enabled the study of cell-population kinetics of infection and cell death by DENV-2, ZIKV, and yellow fever virus. We anticipate that future studies of viral infection kinetics with this reporter system will enable basic investigations of virus-host interactions and facilitate future applications in antiviral drug research to manage flavivirus infections.
Keywords: NS2B–NS3; Zika virus (ZIKV); cell death; cell-based reporter; dengue virus (DENV); flavivirus; fluorescence; internal cleavage; plaque assay; protease.
© 2020 Arias-Arias et al.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article
Figures

Similar articles
-
Generation and Implementation of Reporter BHK-21 Cells for Live Imaging of Flavivirus Infection.Bio Protoc. 2021 Mar 5;11(5):e3942. doi: 10.21769/BioProtoc.3942. eCollection 2021 Mar 5. Bio Protoc. 2021. PMID: 33796616 Free PMC article.
-
Potential Role of Flavivirus NS2B-NS3 Proteases in Viral Pathogenesis and Anti-flavivirus Drug Discovery Employing Animal Cells and Models: A Review.Viruses. 2021 Dec 28;14(1):44. doi: 10.3390/v14010044. Viruses. 2021. PMID: 35062249 Free PMC article. Review.
-
Zika Virus NS2A-Mediated Virion Assembly.mBio. 2019 Oct 29;10(5):e02375-19. doi: 10.1128/mBio.02375-19. mBio. 2019. PMID: 31662457 Free PMC article.
-
Dynamic Interactions of Post Cleaved NS2B Cofactor and NS3 Protease Identified by Integrative Structural Approaches.Viruses. 2022 Jun 30;14(7):1440. doi: 10.3390/v14071440. Viruses. 2022. PMID: 35891424 Free PMC article.
-
The Inhibition of NS2B/NS3 Protease: A New Therapeutic Opportunity to Treat Dengue and Zika Virus Infection.Int J Mol Sci. 2024 Apr 16;25(8):4376. doi: 10.3390/ijms25084376. Int J Mol Sci. 2024. PMID: 38673962 Free PMC article. Review.
Cited by
-
Generation and Implementation of Reporter BHK-21 Cells for Live Imaging of Flavivirus Infection.Bio Protoc. 2021 Mar 5;11(5):e3942. doi: 10.21769/BioProtoc.3942. eCollection 2021 Mar 5. Bio Protoc. 2021. PMID: 33796616 Free PMC article.
-
The Virus-Induced Cytopathic Effect.Subcell Biochem. 2023;106:197-210. doi: 10.1007/978-3-031-40086-5_7. Subcell Biochem. 2023. PMID: 38159228
-
Protein Abundance of Drug Transporters in Human Hepatitis C Livers.Int J Mol Sci. 2022 Jul 19;23(14):7947. doi: 10.3390/ijms23147947. Int J Mol Sci. 2022. PMID: 35887291 Free PMC article.
-
A Versatile Reporter System To Monitor Virus-Infected Cells and Its Application to Dengue Virus and SARS-CoV-2.J Virol. 2021 Jan 28;95(4):e01715-20. doi: 10.1128/JVI.01715-20. Print 2021 Jan 28. J Virol. 2021. PMID: 33257477 Free PMC article.
-
Evaluating the specificity of flavivirus proteases in Aedes aegypti cells for dengue virus 2-derived cleavage sites.PLoS One. 2024 Dec 3;19(12):e0309095. doi: 10.1371/journal.pone.0309095. eCollection 2024. PLoS One. 2024. PMID: 39625906 Free PMC article.
References
-
- Lindenbach B., Murray C. L., Thiel H.-J., and Rice C. M. (2013) Flaviviridae. In Fields Virology, pp. 712–746, Lippincott Williams & Wilkins, Philadelphia, Pennsylvania
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

