LncRNA ZFPM2-AS1 promotes lung adenocarcinoma progression by interacting with UPF1 to destabilize ZFPM2
- PMID: 31919993
- PMCID: PMC7191191
- DOI: 10.1002/1878-0261.12631
LncRNA ZFPM2-AS1 promotes lung adenocarcinoma progression by interacting with UPF1 to destabilize ZFPM2
Abstract
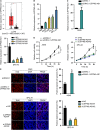
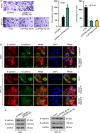
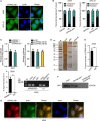
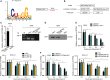
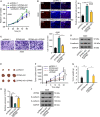
Lung adenocarcinoma (LUAD), a histological subclass of non-small-cell lung cancer, is globally the leading cause of cancer-related deaths. Long noncoding RNAs (lncRNAs) are emerging as cancer regulators. Zinc finger protein multitype 2 antisense RNA 1 (ZFPM2-AS1) is an oncogene in gastric cancer, but its functions have not been investigated in LUAD. We showed that ZFPM2-AS1 expression is high in LUAD samples based on GEPIA database (http://gepia.cancer-pku.cn/) and validated ZFPM2-AS1 upregulation in LUAD cell lines. Functionally, ZFPM2-AS1 facilitated proliferation, invasion, and epithelial-to-mesenchymal transition of LUAD cells. Thereafter, we found that ZFPM2 was negatively regulated by ZFPM2-AS1, and identified the suppressive effect of ZFPM2 regulation by ZFPM2-AS1 on LUAD progression. Mechanistically, we showed that ZFPM2-AS1 interacted with up-frameshift 1 (UPF1) to regulate mRNA decay of ZFPM2. Rescue assays in vitro and in vivo confirmed that ZFPM2-AS1 regulated LUAD progression and tumor growth through ZFPM2. Taken together, our findings demonstrate a role for the ZFPM2-AS1-UPF1-ZFPM2 axis in LUAD progression, suggesting ZFPM2-AS1 as a new potential target for LUAD treatment.
Keywords: UPF1; ZFPM2; ZFPM2-AS1; lung adenocarcinoma; mRNA decay.
© 2020 The Authors. Published by FEBS Press and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
lncRNA ZFPM2-AS1 promotes proliferation via miR-18b-5p/VMA21 axis in lung adenocarcinoma.J Cell Biochem. 2020 Jan;121(1):313-321. doi: 10.1002/jcb.29176. Epub 2019 Jul 11. J Cell Biochem. 2020. PMID: 31297866
-
Long noncoding RNA ZFPM2-AS1 is involved in lung adenocarcinoma via miR-511-3p/AFF4 pathway.J Cell Biochem. 2020 Mar;121(3):2534-2542. doi: 10.1002/jcb.29476. Epub 2019 Nov 6. J Cell Biochem. 2020. Retraction in: J Cell Biochem. 2022 Nov;123(11):1874. doi: 10.1002/jcb.30337. PMID: 31692047 Retracted.
-
LncRNA PCBP1-AS1 correlated with the functional states of cancer cells and inhibited lung adenocarcinoma metastasis by suppressing the EMT progression.Carcinogenesis. 2021 Jul 16;42(7):931-939. doi: 10.1093/carcin/bgab047. Carcinogenesis. 2021. PMID: 34107009
-
Long Non-coding RNA ZFPM2-AS1: A Novel Biomarker in the Pathogenesis of Human Cancers.Mol Biotechnol. 2022 Jul;64(7):725-742. doi: 10.1007/s12033-021-00443-3. Epub 2022 Jan 31. Mol Biotechnol. 2022. PMID: 35098483 Review.
-
The Emerging Role of Thymopoietin-Antisense RNA 1 as Long Noncoding RNA in the Pathogenesis of Human Cancers.DNA Cell Biol. 2021 Jul;40(7):848-857. doi: 10.1089/dna.2021.0024. Epub 2021 Jun 4. DNA Cell Biol. 2021. PMID: 34096793 Review.
Cited by
-
Identification and Validation of UPF1 as a Novel Prognostic Biomarker in Renal Clear Cell Carcinoma.Genes (Basel). 2022 Nov 20;13(11):2166. doi: 10.3390/genes13112166. Genes (Basel). 2022. PMID: 36421841 Free PMC article.
-
UPF1-From mRNA Degradation to Human Disorders.Cells. 2023 Jan 27;12(3):419. doi: 10.3390/cells12030419. Cells. 2023. PMID: 36766761 Free PMC article. Review.
-
Comprehensive analysis of ZFPM2-AS1 prognostic value, immune microenvironment, drug sensitivity, and co-expression network: from gastric adenocarcinoma to pan-cancers.Discov Oncol. 2022 Apr 13;13(1):24. doi: 10.1007/s12672-022-00487-0. Discov Oncol. 2022. PMID: 35416526 Free PMC article.
-
BMSC-Derived Exosomes Carrying lncRNA-ZFAS1 Alleviate Pulmonary Ischemia/Reperfusion Injury by UPF1-Mediated mRNA Decay of FOXD1.Mol Neurobiol. 2023 May;60(5):2379-2396. doi: 10.1007/s12035-022-03129-2. Epub 2023 Jan 18. Mol Neurobiol. 2023. PMID: 36652050
-
Comprehensive landscape and future perspective of long noncoding RNAs in non-small cell lung cancer: it takes a village.Mol Ther. 2023 Dec 6;31(12):3389-3413. doi: 10.1016/j.ymthe.2023.09.015. Epub 2023 Sep 21. Mol Ther. 2023. PMID: 37740493 Free PMC article. Review.
References
-
- Azzalin CM and Lingner J (2006) The Human RNA surveillance factor UPF1 Is required for S phase progression and genome stability. Curr Biol 16, 433–439. - PubMed
-
- He J, Zuo Q, Hu B, Jin H, Wang C, Cheng Z, Deng X, Yang C, Ruan H, Yu C et al (2019) A novel, liver‐specific long noncoding RNA LINC01093 suppresses HCC progression by interaction with IGF2BP1 to facilitate decay of GLI1 mRNA. Cancer Lett 450, 98–109. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

