Structural Insights Into TDP-43 and Effects of Post-translational Modifications
- PMID: 31920533
- PMCID: PMC6934062
- DOI: 10.3389/fnmol.2019.00301
Structural Insights Into TDP-43 and Effects of Post-translational Modifications
Erratum in
-
Corrigendum: Structural Insights Into TDP-43 and Effects of Post-translational Modifications.Front Mol Neurosci. 2020 Apr 2;13:45. doi: 10.3389/fnmol.2020.00045. eCollection 2020. Front Mol Neurosci. 2020. PMID: 32300293 Free PMC article.
Abstract
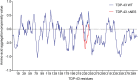
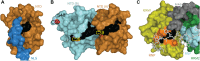
Transactive response DNA binding protein (TDP-43) is a key player in neurodegenerative diseases. In this review, we have gathered and presented structural information on the different regions of TDP-43 with high resolution structures available. A thorough understanding of TDP-43 structure, effect of modifications, aggregation and sites of localization is necessary as we develop therapeutic strategies targeting TDP-43 for neurodegenerative diseases. We discuss how different domains as well as post-translational modification may influence TDP-43 overall structure, aggregation and droplet formation. The primary aim of the review is to utilize structural insights as we develop an understanding of the deleterious behavior of TDP-43 and highlight locations of established and proposed post-translation modifications. TDP-43 structure and effect on localization is paralleled by many RNA-binding proteins and this review serves as an example of how structure may be modulated by numerous compounding elements.
Keywords: RRM domain; TDP-43 = TAR DNA–binding protein 43; post-translational modification; structure; subdomains.
Copyright © 2019 François-Moutal, Perez-Miller, Scott, Miranda, Mollasalehi and Khanna.
Figures

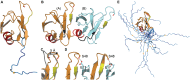
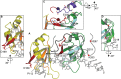





References
-
- Arai T., Hasegawa M., Akiyama H., Ikeda K., Nonaka T., Mori H., et al. (2006). TDP-43 is a component of ubiquitin-positive tau-negative inclusions in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Biochem. Biophys. Res. Commun. 351 602–611. 10.1016/j.bbrc.2006.10.093 - DOI - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

