Differences in the chitinolytic activity of mammalian chitinases on soluble and insoluble substrates
- PMID: 31930591
- PMCID: PMC7096708
- DOI: 10.1002/pro.3822
Differences in the chitinolytic activity of mammalian chitinases on soluble and insoluble substrates
Abstract
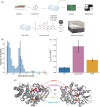
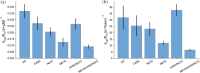
Chitin is an abundant polysaccharide used by many organisms for structural rigidity and water repulsion. As such, the insoluble crystalline structure of chitin poses significant challenges for enzymatic degradation. Acidic mammalian chitinase, a processive glycosyl hydrolase, is the primary enzyme involved in the degradation of environmental chitin in mammalian lungs. Mutations to acidic mammalian chitinase have been associated with asthma, and genetic deletion in mice increases morbidity and mortality with age. We initially set out to reverse this phenotype by engineering hyperactive acidic mammalian chitinase variants. Using a screening approach with commercial fluorogenic substrates, we identified mutations with consistent increases in activity. To determine whether the activity increases observed were consistent with more biologically relevant chitin substrates, we developed new assays to quantify chitinase activity with insoluble chitin, and identified a one-pot fluorogenic assay that is sufficiently sensitive to quantify changes to activity due to the addition or removal of a carbohydrate-binding domain. We show that the activity increases from our directed evolution screen were lost when insoluble substrates were used. In contrast, naturally occurring gain-of-function mutations gave similar results with oligomeric and insoluble substrates. We also show that activity differences between acidic mammalian chitinase and chitotriosidase are reduced with insoluble substrate, suggesting that previously reported activity differences with oligomeric substrates may have been driven by differential substrate specificity. These results highlight the need for assays against physiological substrates when engineering metabolic enzymes, and provide a new one-pot assay that may prove to be broadly applicable to engineering glycosyl hydrolases.
Keywords: chitin; directed evolution; enzymes; glycobiology.
© 2020 The Authors. Protein Science published by Wiley Periodicals, Inc. on behalf of The Protein Society.
Conflict of interest statement
BAB and JSF are inventors on a provisional patent application for the mutants described herein and their use in treating fibrotic lung disease. SJVD and RML are inventors on a pending patent application on the use of chitinases for treating fibrotic lung disease.
Figures



References
-
- Zhu KY, Merzendorfer H, Zhang W, Zhang J, Muthukrishnan S. Biosynthesis, turnover, and functions of chitin in insects. Annu Rev Entomol. 2016;61:177–196. - PubMed
-
- Boot RG, Blommaart EF, Swart E, et al. Identification of a novel acidic mammalian chitinase distinct from chitotriosidase. J Biol Chem. 2001;276:6770–6778. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

