Role of circular RNAs in brain development and CNS diseases
- PMID: 31931031
- PMCID: PMC7024016
- DOI: 10.1016/j.pneurobio.2020.101746
Role of circular RNAs in brain development and CNS diseases
Abstract
In mammals, many classes of noncoding RNAs (ncRNAs) are expressed at a much higher level in the brain than in other organs. Recent studies have identified a new class of ncRNAs called circular RNAs (circRNAs), which are produced by back-splicing and fusion of either exons, introns, or both exon-intron into covalently closed loops. The circRNAs are also highly enriched in the brain and increase continuously from the embryonic to the adult stage. Although the functional significance and mechanism of action of circRNAs are still being actively explored, they are thought to regulate the transcription of their host genes and sequestration of miRNAs and RNA binding proteins. Some circRNAs are also shown to have translation potential to form peptides. The expression and abundance of circRNAs seem to be spatiotemporally maintained in a normal brain. Altered expression of circRNAs is also thought to mediate several disorders, including brain-tumor growth, and acute and chronic neurodegenerative disorders by affecting mechanisms such as angiogenesis, neuronal plasticity, autophagy, apoptosis, and inflammation. This review discusses the involvement of various circRNAs in brain development and CNS diseases. A better understanding of the circRNA function will help to develop novel therapeutic strategies to treat CNS complications.
Keywords: Acute and chronic neurodegeneration; Brain; Cancer; CircRNAs; Development; miRNAs.
Copyright © 2020 Elsevier Ltd. All rights reserved.
Conflict of interest statement
Declaration of Competing Interest Authors declare no conflict of interest.
Figures
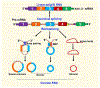



References
-
- Aktas T, Avsar Ilik I, Maticzka D, Bhardwaj V, Pessoa Rodrigues C, Mittler G, Manke T, Backofen R, Akhtar A, 2017. DHX9 suppresses RNA processing defects originating from the Alu invasion of the human genome. Nature 544, 115119. - PubMed
-
- Arnberg AC, Van Ommen GJ, Grivell LA, Van Bruggen EF, Borst P, 1980. Some yeast mitochondrial RNAs are circular. Cell 19, 313–319. - PubMed
-
- Ashwal-Fluss R, Meyer M, Pamudurti NR, Ivanov A, Bartok O, Hanan M, Evantal N, Memczak S, Rajewsky N, Kadener S, 2014. circRNA biogenesis competes with pre-mRNA splicing. Mol Cell 56, 55–66. - PubMed
-
- Bai Y, Zhang Y, Han B, Yang L, Chen X, Huang R, Wu F, Chao J, Liu P, Hu G, Zhang JH, Yao H, 2018. Circular RNA DLGAP4 Ameliorates Ischemic Stroke Outcomes by Targeting miR-143 to Regulate Endothelial-Mesenchymal Transition Associated with Blood-Brain Barrier Integrity. J Neurosci 38, 32–50. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

