Cas9/gRNA-mediated genome editing of yeast mitochondria and Chlamydomonas chloroplasts
- PMID: 31934513
- PMCID: PMC6951285
- DOI: 10.7717/peerj.8362
Cas9/gRNA-mediated genome editing of yeast mitochondria and Chlamydomonas chloroplasts
Abstract
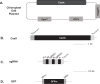
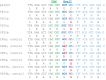
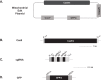
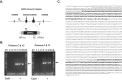
We present a new approach to edit both mitochondrial and chloroplast genomes. Organelles have been considered off-limits to CRISPR due to their impermeability to most RNA and DNA. This has prevented applications of Cas9/gRNA-mediated genome editing in organelles while the tool has been widely used for engineering of nuclear DNA in a number of organisms in the last several years. To overcome the hurdle, we designed a new approach to enable organelle genome editing. The plasmids, designated "Edit Plasmids," were constructed with two expression cassettes, one for the expression of Cas9, codon-optimized for each organelle, under promoters specific to each organelle, and the other cassette for the expression of guide RNAs under another set of promoters specific to each organelle. In addition, Edit Plasmids were designed to carry the donor DNA for integration between two double-strand break sites induced by Cas9/gRNAs. Each donor DNA was flanked by the regions homologous to both ends of the integration site that were short enough to minimize spontaneous recombination events. Furthermore, the donor DNA was so modified that it did not carry functional gRNA target sites, allowing the stability of the integrated DNA without being excised by further Cas9/gRNAs activity. Edit Plasmids were introduced into organelles through microprojectile transformation. We confirmed donor DNA insertion at the target sites facilitated by homologous recombination only in the presence of Cas9/gRNA activity in yeast mitochondria and Chlamydomonas chloroplasts. We also showed that Edit Plasmids persist and replicate in mitochondria autonomously for several dozens of generations in the presence of the wild-type genomes. Finally, we did not find insertions and/or deletions at one of the Cas9 cleavage sites in Chloroplasts, which are otherwise hallmarks of Cas9/gRNA-mediated non-homologous end joining (NHEJ) repair events in nuclear DNA. This is consistent with previous reports of the lack of NHEJ repair system in most bacteria, which are believed to be ancestors of organelles. This is the first demonstration of CRISPR-mediated genome editing in both mitochondria and chloroplasts in two distantly related organisms. The Edit Plasmid approach is expected to open the door to engineer organelle genomes of a wide range of organisms in a precise fashion.
Keywords: CRISPR; Cas9/gRNA; Chlamydomonas; Chloroplast; DNA replacement; Edit Plasmid; Genome editing; Mitochondria; Organelle; Yeast.
© 2020 Yoo et al.
Conflict of interest statement
Hajime Sakai is CEO at Napigen. Byung-Chun Yoo, Narendra Yadav and Emil Orozco, Jr are employees at Napigen. All authors are co-inventors on patent application US2019/0136249 A1 “organelle genome modification using polynucleotide guided endonuclease,” submitted by Napigen. The authors declare that they have no other competing interests.
Figures


Similar articles
-
Harnessing accurate non-homologous end joining for efficient precise deletion in CRISPR/Cas9-mediated genome editing.Genome Biol. 2018 Oct 19;19(1):170. doi: 10.1186/s13059-018-1518-x. Genome Biol. 2018. PMID: 30340517 Free PMC article.
-
Two Distinct Approaches for CRISPR-Cas9-Mediated Gene Editing in Cryptococcus neoformans and Related Species.mSphere. 2018 Jun 13;3(3):e00208-18. doi: 10.1128/mSphereDirect.00208-18. Print 2018 Jun 27. mSphere. 2018. PMID: 29898980 Free PMC article.
-
CRISPR-Cas9-Mediated Genome Editing in Leishmania donovani.mBio. 2015 Jul 21;6(4):e00861. doi: 10.1128/mBio.00861-15. mBio. 2015. PMID: 26199327 Free PMC article.
-
Gene Editing in Clinical Practice: Where are We?Indian J Clin Biochem. 2019 Jan;34(1):19-25. doi: 10.1007/s12291-018-0804-4. Epub 2019 Jan 1. Indian J Clin Biochem. 2019. PMID: 30728669 Free PMC article. Review.
-
The application of CRISPR-Cas9 genome editing in Caenorhabditis elegans.J Genet Genomics. 2015 Aug 20;42(8):413-21. doi: 10.1016/j.jgg.2015.06.005. Epub 2015 Jun 26. J Genet Genomics. 2015. PMID: 26336798 Free PMC article. Review.
Cited by
-
The Mitochondrial Genome in Aging and Disease and the Future of Mitochondrial Therapeutics.Biomedicines. 2022 Feb 18;10(2):490. doi: 10.3390/biomedicines10020490. Biomedicines. 2022. PMID: 35203698 Free PMC article. Review.
-
Mechanism and Utilization of Ogura Cytoplasmic Male Sterility in Cruciferae Crops.Int J Mol Sci. 2022 Aug 13;23(16):9099. doi: 10.3390/ijms23169099. Int J Mol Sci. 2022. PMID: 36012365 Free PMC article. Review.
-
Strategies for mitochondrial gene editing.Comput Struct Biotechnol J. 2021 Jun 4;19:3319-3329. doi: 10.1016/j.csbj.2021.06.003. eCollection 2021. Comput Struct Biotechnol J. 2021. PMID: 34188780 Free PMC article. Review.
-
Green giant-a tiny chloroplast genome with mighty power to produce high-value proteins: history and phylogeny.Plant Biotechnol J. 2021 Mar;19(3):430-447. doi: 10.1111/pbi.13556. Epub 2021 Feb 22. Plant Biotechnol J. 2021. PMID: 33484606 Free PMC article. Review.
-
TALE-based organellar genome editing and gene expression in plants.Plant Cell Rep. 2024 Feb 10;43(3):61. doi: 10.1007/s00299-024-03150-w. Plant Cell Rep. 2024. PMID: 38336900 Review.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources

