Epidermal autonomous VEGFA/Flt1/Nrp1 functions mediate psoriasis-like disease
- PMID: 31934626
- PMCID: PMC6949033
- DOI: 10.1126/sciadv.aax5849
Epidermal autonomous VEGFA/Flt1/Nrp1 functions mediate psoriasis-like disease
Abstract
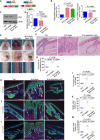
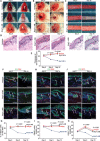
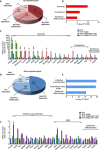
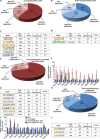
Psoriasis is a common chronic skin disorder characterized by keratinocyte hyperproliferation with altered differentiation accompanied by inflammation and increased angiogenesis. It remains unclear whether the first events that initiate psoriasis development occur in keratinocytes or inflammatory cells. Here, using different psoriasis mouse models, we showed that conditional deletion of Flt1 or Nrp1 in epidermal cells inhibited psoriasis mediated by Vegfa overexpression or c-Jun/JunB deletion. Administration of anti-Nrp1 antibody reverted the psoriasis phenotype. Using transcriptional and chromatin profiling of epidermal cells following Vegfa overexpression together with Flt1 or Nrp1 deletion, we identified the gene regulatory network regulated by Vegfa/Nrp1/Flt1 during psoriasis development and uncovered a key role of Fosl1 in regulating the chromatin remodeling mediated by Vegfa overexpression in keratinocytes. In conclusion, our study identifies an epidermal autonomous function of Vegfa/Nrp1/Flt1 that mediates psoriatic-like disease and demonstrates the clinical relevance of blocking Vegfa/Nrp1/Flt1 axis in psoriasis.
Copyright © 2020 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works. Distributed under a Creative Commons Attribution NonCommercial License 4.0 (CC BY-NC).
Figures

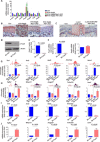
Similar articles
-
Epidermal expression of neuropilin 1 protects murine keratinocytes from UVB-induced apoptosis.PLoS One. 2012;7(12):e50944. doi: 10.1371/journal.pone.0050944. Epub 2012 Dec 10. PLoS One. 2012. PMID: 23251405 Free PMC article.
-
Differential dependency of human glioblastoma cells on vascular endothelial growth factor‑A signaling via neuropilin‑1.Int J Oncol. 2022 Oct;61(4):122. doi: 10.3892/ijo.2022.5412. Epub 2022 Aug 31. Int J Oncol. 2022. PMID: 36043525 Free PMC article.
-
Neuropilin-1 controls vascular permeability through juxtacrine regulation of endothelial adherens junctions.Angiogenesis. 2024 Dec 12;28(1):7. doi: 10.1007/s10456-024-09963-3. Angiogenesis. 2024. PMID: 39668325 Free PMC article.
-
Jun signalling in the epidermis: From developmental defects to psoriasis and skin tumors.Int J Biochem Cell Biol. 2006;38(7):1043-9. doi: 10.1016/j.biocel.2005.11.011. Epub 2005 Dec 20. Int J Biochem Cell Biol. 2006. PMID: 16423552 Review.
-
Neuropilin-1 enforces extracellular matrix signalling via ABL1 to promote angiogenesis.Biochem Soc Trans. 2014 Oct;42(5):1429-34. doi: 10.1042/BST20140141. Biochem Soc Trans. 2014. PMID: 25233427 Review.
Cited by
-
Sclerotic-Type Cutaneous Chronic Graft-Versus-Host Disease Exhibits Activation of T Helper 1 and OX40 Cytokines.J Invest Dermatol. 2024 Mar;144(3):563-572.e9. doi: 10.1016/j.jid.2023.08.026. Epub 2023 Sep 23. J Invest Dermatol. 2024. PMID: 37742913 Free PMC article.
-
Rutin ameliorates imiquimod-induced psoriasis-like skin lesions by inhibiting oxidative stress injury and the inflammatory response in mice via the Keap1/Nrf2 signaling pathway.Sci Rep. 2025 Jul 1;15(1):20712. doi: 10.1038/s41598-025-08451-y. Sci Rep. 2025. PMID: 40596643 Free PMC article.
-
Pan-Cancer Analysis of BUB1B/hsa-miR-130a-3p Axis and Identification of Circulating hsa-miR-130a-3p as a Potential Biomarker for Cancer Risk Assessment.Evid Based Complement Alternat Med. 2022 Sep 22;2022:3261300. doi: 10.1155/2022/3261300. eCollection 2022. Evid Based Complement Alternat Med. 2022. PMID: 36185088 Free PMC article.
-
Vascular Endothelial Growth Factor a Promotes Chronic Itch via VEGFA-VEGFR2-PI3K-TRPV1 Axis in Allergic Contact Dermatitis.J Inflamm Res. 2024 Oct 17;17:7423-7439. doi: 10.2147/JIR.S470094. eCollection 2024. J Inflamm Res. 2024. PMID: 39435259 Free PMC article.
-
Keratinocyte-derived VEGF-A is an essential pro-migratory autocrine mediator, acting through the KDR/GEF-H1/RhoA pathway.Front Cell Dev Biol. 2025 Jul 17;13:1601887. doi: 10.3389/fcell.2025.1601887. eCollection 2025. Front Cell Dev Biol. 2025. PMID: 40746859 Free PMC article.
References
-
- Canavese M., Altruda F., Ruzicka T., Schauber J., Vascular endothelial growth factor (VEGF) in the pathogenesis of psoriasis—A possible target for novel therapies? J. Dermatol. Sci. 58, 171–176 (2010). - PubMed
-
- Wang Z., Liang W., Zhang B., Lv M., Wang J., Zhang L., Single nucleotide polymorphisms of VEGF gene and psoriasis risk. J. Dermatol. Sci. 49, 263–265 (2008). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

