Rules and Exceptions: The Role of Chromosomal ParB in DNA Segregation and Other Cellular Processes
- PMID: 31940850
- PMCID: PMC7022226
- DOI: 10.3390/microorganisms8010105
Rules and Exceptions: The Role of Chromosomal ParB in DNA Segregation and Other Cellular Processes
Abstract
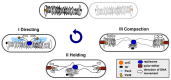
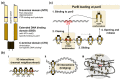
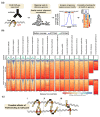
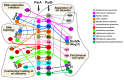
The segregation of newly replicated chromosomes in bacterial cells is a highly coordinated spatiotemporal process. In the majority of bacterial species, a tripartite ParAB-parS system, composed of an ATPase (ParA), a DNA-binding protein (ParB), and its target(s) parS sequence(s), facilitates the initial steps of chromosome partitioning. ParB nucleates around parS(s) located in the vicinity of newly replicated oriCs to form large nucleoprotein complexes, which are subsequently relocated by ParA to distal cellular compartments. In this review, we describe the role of ParB in various processes within bacterial cells, pointing out interspecies differences. We outline recent progress in understanding the ParB nucleoprotein complex formation and its role in DNA segregation, including ori positioning and anchoring, DNA condensation, and loading of the structural maintenance of chromosome (SMC) proteins. The auxiliary roles of ParBs in the control of chromosome replication initiation and cell division, as well as the regulation of gene expression, are discussed. Moreover, we catalog ParB interacting proteins. Overall, this work highlights how different bacterial species adapt the DNA partitioning ParAB-parS system to meet their specific requirements.
Keywords: ParB; cell division; chromosome segregation; gene expression regulation; partitioning proteins; segrosome.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

