The Bcl-2 Family: Ancient Origins, Conserved Structures, and Divergent Mechanisms
- PMID: 31940915
- PMCID: PMC7022251
- DOI: 10.3390/biom10010128
The Bcl-2 Family: Ancient Origins, Conserved Structures, and Divergent Mechanisms
Abstract
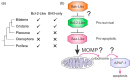
Intrinsic apoptosis, the response to intracellular cell death stimuli, is regulated by the interplay of the B-cell lymphoma 2 (Bcl-2) family and their membrane interactions. Bcl-2 proteins mediate a number of processes including development, homeostasis, autophagy, and innate and adaptive immune responses and their dysregulation underpins a host of diseases including cancer. The Bcl-2 family is characterized by the presence of conserved sequence motifs called Bcl-2 homology motifs, as well as a transmembrane region, which form the interaction sites and intracellular location mechanism, respectively. Bcl-2 proteins have been recognized in the earliest metazoans including Porifera (sponges), Placozoans, and Cnidarians (e.g., Hydra). A number of viruses have gained Bcl-2 homologs and subvert innate immunity and cellular apoptosis for their replication, but they frequently have very different sequences to their host Bcl-2 analogs. Though most mechanisms of apoptosis initiation converge on activation of caspases that destroy the cell from within, the numerous gene insertions, deletions, and duplications during evolution have led to a divergence in mechanisms of intrinsic apoptosis. Currently, the action of the Bcl-2 family is best understood in vertebrates and nematodes but new insights are emerging from evolutionarily earlier organisms. This review focuses on the mechanisms underpinning the activity of Bcl-2 proteins including their structures and interactions, and how they have changed over the course of evolution.
Keywords: Bcl-2; apoptosis; evolution; mechanism; structure analysis.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


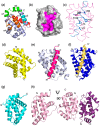
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

