Privacy-preserving model learning on a blockchain network-of-networks
- PMID: 31943009
- PMCID: PMC7025358
- DOI: 10.1093/jamia/ocz214
Privacy-preserving model learning on a blockchain network-of-networks
Abstract
Objective: To facilitate clinical/genomic/biomedical research, constructing generalizable predictive models using cross-institutional methods while protecting privacy is imperative. However, state-of-the-art methods assume a "flattened" topology, while real-world research networks may consist of "network-of-networks" which can imply practical issues including training on small data for rare diseases/conditions, prioritizing locally trained models, and maintaining models for each level of the hierarchy. In this study, we focus on developing a hierarchical approach to inherit the benefits of the privacy-preserving methods, retain the advantages of adopting blockchain, and address practical concerns on a research network-of-networks.
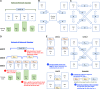
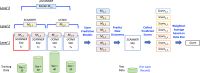
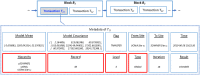
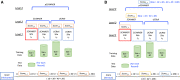
Materials and methods: We propose a framework to combine level-wise model learning, blockchain-based model dissemination, and a novel hierarchical consensus algorithm for model ensemble. We developed an example implementation HierarchicalChain (hierarchical privacy-preserving modeling on blockchain), evaluated it on 3 healthcare/genomic datasets, as well as compared its predictive correctness, learning iteration, and execution time with a state-of-the-art method designed for flattened network topology.
Results: HierarchicalChain improves the predictive correctness for small training datasets and provides comparable correctness results with the competing method with higher learning iteration and similar per-iteration execution time, inherits the benefits of the privacy-preserving learning and advantages of blockchain technology, and immutable records models for each level.
Discussion: HierarchicalChain is independent of the core privacy-preserving learning method, as well as of the underlying blockchain platform. Further studies are warranted for various types of network topology, complex data, and privacy concerns.
Conclusion: We demonstrated the potential of utilizing the information from the hierarchical network-of-networks topology to improve prediction.
Keywords: blockchain distributed ledger technology; clinical information systems; decision support systems; hierarchical network; privacy-preserving predictive modeling.
© The Author(s) 2020. Published by Oxford University Press on behalf of the American Medical Informatics Association.
Figures
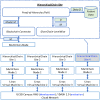


Similar articles
-
Fair compute loads enabled by blockchain: sharing models by alternating client and server roles.J Am Med Inform Assoc. 2019 May 1;26(5):392-403. doi: 10.1093/jamia/ocy180. J Am Med Inform Assoc. 2019. PMID: 30892656 Free PMC article.
-
Quorum-based model learning on a blockchain hierarchical clinical research network using smart contracts.Int J Med Inform. 2023 Jan;169:104924. doi: 10.1016/j.ijmedinf.2022.104924. Epub 2022 Nov 9. Int J Med Inform. 2023. PMID: 36402113 Free PMC article.
-
EXpectation Propagation LOgistic REgRession on permissioned blockCHAIN (ExplorerChain): decentralized online healthcare/genomics predictive model learning.J Am Med Inform Assoc. 2020 May 1;27(5):747-756. doi: 10.1093/jamia/ocaa023. J Am Med Inform Assoc. 2020. PMID: 32364235 Free PMC article.
-
Privacy-Preserving Methods for Feature Engineering Using Blockchain: Review, Evaluation, and Proof of Concept.J Med Internet Res. 2019 Aug 14;21(8):e13600. doi: 10.2196/13600. J Med Internet Res. 2019. PMID: 31414666 Free PMC article. Review.
-
A survey of security, privacy and trust issues in vehicular computation offloading and their solutions using blockchain.Open Res Eur. 2023 Oct 25;3:110. doi: 10.12688/openreseurope.16189.2. eCollection 2023. Open Res Eur. 2023. PMID: 37969246 Free PMC article. Review.
Cited by
-
Distributed cross-learning for equitable federated models - privacy-preserving prediction on data from five California hospitals.Nat Commun. 2025 Feb 5;16(1):1371. doi: 10.1038/s41467-025-56510-9. Nat Commun. 2025. PMID: 39910076 Free PMC article.
-
CertificateChain: decentralized healthcare training certificate management system using blockchain and smart contracts.JAMIA Open. 2022 Mar 14;5(1):ooac019. doi: 10.1093/jamiaopen/ooac019. eCollection 2022 Apr. JAMIA Open. 2022. PMID: 35571362 Free PMC article.
-
Innovation is key for advancing the science of biomedical and health informatics and for publishing in JAMIA.J Am Med Inform Assoc. 2020 Mar 1;27(3):341-342. doi: 10.1093/jamia/ocaa002. J Am Med Inform Assoc. 2020. PMID: 32055863 Free PMC article. No abstract available.
-
Functional genomics data: privacy risk assessment and technological mitigation.Nat Rev Genet. 2022 Apr;23(4):245-258. doi: 10.1038/s41576-021-00428-7. Epub 2021 Nov 10. Nat Rev Genet. 2022. PMID: 34759381 Review.
-
Benchmarking blockchain-based gene-drug interaction data sharing methods: A case study from the iDASH 2019 secure genome analysis competition blockchain track.Int J Med Inform. 2021 Oct;154:104559. doi: 10.1016/j.ijmedinf.2021.104559. Epub 2021 Aug 18. Int J Med Inform. 2021. PMID: 34474309 Free PMC article.
References
-
- Navathe AS, Conway PH.. Optimizing health information technology's role in enabling comparative effectiveness research. Am J Managed Care 2010; 16 (12 Suppl HIT): SP44–7. - PubMed
-
- Wicks P, Vaughan TE, Massagli MP, Heywood J.. Accelerated clinical discovery using self-reported patient data collected online and a patient-matching algorithm. Nat Biotechnol 2011; 29 (5): 411–4. - PubMed
-
- Grossman JM, Kushner KL, November EA, Lthpolicy PC.. Creating Sustainable Local Health Information Exchanges: Can Barriers to Stakeholder Participation Be Overcome? Washington, DC: Center for Studying Health System Change; 2008. - PubMed
-
- ClinVar. https://www.ncbi.nlm.nih.gov/clinvar/. Accessed June 1, 2017.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

