GWAS: Fast-forwarding gene identification and characterization in temperate Cereals: lessons from Barley - A review
- PMID: 31956447
- PMCID: PMC6961222
- DOI: 10.1016/j.jare.2019.10.013
GWAS: Fast-forwarding gene identification and characterization in temperate Cereals: lessons from Barley - A review
Abstract
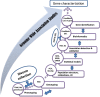
Understanding the genetic complexity of traits is an important objective of small grain temperate cereals yield and adaptation improvements. Bi-parental quantitative trait loci (QTL) linkage mapping is a powerful method to identify genetic regions that co-segregate in the trait of interest within the research population. However, recently, association or linkage disequilibrium (LD) mapping using a genome-wide association study (GWAS) became an approach for unraveling the molecular genetic basis underlying the natural phenotypic variation. Many causative allele(s)/loci have been identified using the power of this approach which had not been detected in QTL mapping populations. In barley (Hordeum vulgare L.), GWAS has been successfully applied to define the causative allele(s)/loci which can be used in the breeding crop for adaptation and yield improvement. This promising approach represents a tremendous step forward in genetic analysis and undoubtedly proved it is a valuable tool in the identification of candidate genes. In this review, we describe the recently used approach for genetic analyses (linkage mapping or association mapping), and then provide the basic genetic and statistical concepts of GWAS, and subsequently highlight the genetic discoveries using GWAS. The review explained how the candidate gene(s) can be detected using state-of-art bioinformatic tools.
Keywords: Association mapping; Barley breeding, GWAS; Gene identification; Hordeum vulgare L; QTL mapping.
© 2019 THE AUTHORS. Published by Elsevier BV on behalf of Cairo University.
Figures


References
-
- Badr A., Muller K., Schafer-Pregl R., El Rabey H., Effgen S., Ibrahim H.H. On the origin and domestication history of Barley (Hordeum vulgare) Mol Biol Evol. 2000;17(4):499–510. - PubMed
-
- Mitchell-Olds T., Willis J.H., Goldstein D.B. Which evolutionary processes influence natural genetic variation for phenotypic traits? Nat Rev Genet. 2007;8(11):845–856. - PubMed
-
- Pourkheirandish M., Hensel G., Kilian B., Senthil N., Chen G., Sameri M. Evolution of the Grain Dispersal System in Barley. Cell. 2015;162(3):527–539. - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Research Materials

