Development of a Biosensor for Detection of Benzoic Acid Derivatives in Saccharomyces cerevisiae
- PMID: 31970152
- PMCID: PMC6959289
- DOI: 10.3389/fbioe.2019.00372
Development of a Biosensor for Detection of Benzoic Acid Derivatives in Saccharomyces cerevisiae
Abstract
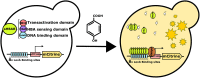
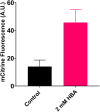
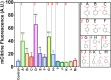
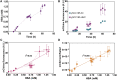
4-hydroxybenzoic acid (pHBA) is an important industrial precursor of muconic acid and liquid crystal polymers whose production is based on the petrochemical industry. In order to decrease our dependency on fossil fuels and improve sustainability, microbial engineering is a particularly appealing approach for replacing traditional chemical techniques. The optimization of microbial strains, however, is still highly constrained by the screening stage. Biosensors have helped to alleviate this problem by decreasing the screening time as well as enabling higher throughput. In this paper, we constructed a synthetic biosensor, named sBAD, consisting of a fusion of the pHBA-binding domain of HbaR from R. palustris, the LexA DNA binding domain at the N-terminus and the transactivation domain B112 at the C-terminus. The response of sBAD was tested in the presence of different benzoic acid derivatives, with cell fluorescence output measured by flow cytometry. The biosensor was found to be activated by the external addition of pHBA in the culture medium, in addition to other carboxylic acids including p-aminobenzoic acid (pABA), salicylic acid, anthranilic acid, aspirin, and benzoic acid. Furthermore, we were able to show that this biosensor could detect the in vivo production of pHBA in a genetically modified yeast strain. A good linearity was observed between the biosensor fluorescence and pHBA concentration. Thus, this biosensor would be well-suited as a high throughput screening tool to produce, via metabolic engineering, benzoic acid derivatives.
Keywords: biosensor; p-aminobenzoic acid; p-hydroxybenzoic acid; synthetic biology; yeast.
Copyright © 2020 Castaño-Cerezo, Fournié, Urban, Faulon and Truan.
Figures


References
-
- Ando S., Arai I., Kiyoto K., Hanai S. (1986). Identification of aromatic monomers in steam-exploded poplar and their influences on ethanol fermentation by Saccharomyces cerevisiae. J. Ferment. Technol. 64, 567–570. 10.1016/0385-6380(86)90084-1 - DOI
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

