All of gene expression (AOE): An integrated index for public gene expression databases
- PMID: 31978081
- PMCID: PMC6980531
- DOI: 10.1371/journal.pone.0227076
All of gene expression (AOE): An integrated index for public gene expression databases
Abstract
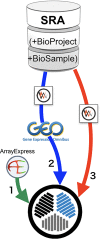
Gene expression data have been archived as microarray and RNA-seq datasets in two public databases, Gene Expression Omnibus (GEO) and ArrayExpress (AE). In 2018, the DNA DataBank of Japan started a similar repository called the Genomic Expression Archive (GEA). These databases are useful resources for the functional interpretation of genes, but have been separately maintained and may lack RNA-seq data, while the original sequence data are available in the Sequence Read Archive (SRA). We constructed an index for those gene expression data repositories, called All Of gene Expression (AOE), to integrate publicly available gene expression data. The web interface of AOE can graphically query data in addition to the application programming interface. By collecting gene expression data from RNA-seq in the SRA, AOE also includes data not included in GEO and AE. AOE is accessible as a search tool from the GEA website and is freely available at https://aoe.dbcls.jp/.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
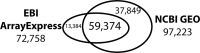

Similar articles
-
GEM-TREND: a web tool for gene expression data mining toward relevant network discovery.BMC Genomics. 2009 Sep 3;10:411. doi: 10.1186/1471-2164-10-411. BMC Genomics. 2009. PMID: 19728865 Free PMC article.
-
GEOmetadb: powerful alternative search engine for the Gene Expression Omnibus.Bioinformatics. 2008 Dec 1;24(23):2798-800. doi: 10.1093/bioinformatics/btn520. Epub 2008 Oct 7. Bioinformatics. 2008. PMID: 18842599 Free PMC article.
-
Microarray retriever: a web-based tool for searching and large scale retrieval of public microarray data.Nucleic Acids Res. 2008 Jul 1;36(Web Server issue):W327-31. doi: 10.1093/nar/gkn213. Epub 2008 May 7. Nucleic Acids Res. 2008. PMID: 18463138 Free PMC article.
-
Gene expression omnibus: microarray data storage, submission, retrieval, and analysis.Methods Enzymol. 2006;411:352-69. doi: 10.1016/S0076-6879(06)11019-8. Methods Enzymol. 2006. PMID: 16939800 Free PMC article. Review.
-
Data storage and analysis in ArrayExpress.Methods Enzymol. 2006;411:370-86. doi: 10.1016/S0076-6879(06)11020-4. Methods Enzymol. 2006. PMID: 16939801 Review.
Cited by
-
Comparison of Oxidative and Hypoxic Stress Responsive Genes from Meta-Analysis of Public Transcriptomes.Biomedicines. 2021 Dec 3;9(12):1830. doi: 10.3390/biomedicines9121830. Biomedicines. 2021. PMID: 34944646 Free PMC article.
-
Meta-Analysis of Public RNA Sequencing Data Revealed Potential Key Genes Associated with Reproductive Division of Labor in Social Hymenoptera and Termites.Int J Mol Sci. 2023 May 6;24(9):8353. doi: 10.3390/ijms24098353. Int J Mol Sci. 2023. PMID: 37176060 Free PMC article.
-
Meta-Analysis of Hypoxic Transcriptomes from Public Databases.Biomedicines. 2020 Jan 9;8(1):10. doi: 10.3390/biomedicines8010010. Biomedicines. 2020. PMID: 31936636 Free PMC article.
-
Integrating Statistical and Machine-Learning Approach for Meta-Analysis of Bisphenol A-Exposure Datasets Reveals Effects on Mouse Gene Expression within Pathways of Apoptosis and Cell Survival.Int J Mol Sci. 2021 Oct 5;22(19):10785. doi: 10.3390/ijms221910785. Int J Mol Sci. 2021. PMID: 34639124 Free PMC article.
-
Multi-Omic Meta-Analysis of Transcriptomes and the Bibliome Uncovers Novel Hypoxia-Inducible Genes.Biomedicines. 2021 May 20;9(5):582. doi: 10.3390/biomedicines9050582. Biomedicines. 2021. PMID: 34065451 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

