The Autophagy Regulatory Molecule CSRP3 Interacts with LC3 and Protects Against Muscular Dystrophy
- PMID: 31979369
- PMCID: PMC7037376
- DOI: 10.3390/ijms21030749
The Autophagy Regulatory Molecule CSRP3 Interacts with LC3 and Protects Against Muscular Dystrophy
Abstract
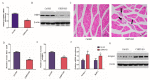
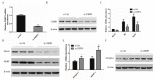
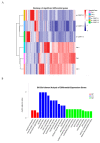
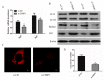
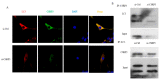
CSRP3/MLP (cysteine-rich protein 3/muscle Lim protein), a member of the cysteine-rich protein family, is a muscle-specific LIM-only factor specifically expressed in skeletal muscle. CSRP3 is critical in maintaining the structure and function of normal muscle. To investigate the mechanism of disease in CSRP3 myopathy, we performed siRNA-mediated CSRP3 knockdown in chicken primary myoblasts. CSRP3 silencing resulted in the down-regulation of the expression of myogenic genes and the up-regulation of atrophy-related gene expressions. We found that CSRP3 interacted with LC3 protein to promote the formation of autophagosomes during autophagy. CSRP3-silencing impaired myoblast autophagy, as evidenced by inhibited autophagy-related ATG5 and ATG7 mRNA expression levels, and inhibited LC3II and Beclin-1 protein accumulation. In addition, impaired autophagy in CSRP3-silenced cells resulted in increased sensitivity to apoptosis cell death. CSRP3-silenced cells also showed increased caspase-3 and caspase-9 cleavage. Moreover, apoptosis induced by CSRP3 silencing was alleviated after autophagy activation. Together, these results indicate that CSRP3 promotes the correct formation of autophagosomes through its interaction with LC3 protein, which has an important role in skeletal muscle remodeling and maintenance.
Keywords: CSRP3; apoptosis; atrophy; autophagy; myoblasts.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
FHL1 regulates myoblast differentiation and autophagy through its interaction with LC3.J Cell Physiol. 2020 May;235(5):4667-4678. doi: 10.1002/jcp.29345. Epub 2019 Oct 21. J Cell Physiol. 2020. PMID: 31637727
-
Increase in HDAC9 suppresses myoblast differentiation via epigenetic regulation of autophagy in hypoxia.Cell Death Dis. 2019 Jul 18;10(8):552. doi: 10.1038/s41419-019-1763-2. Cell Death Dis. 2019. PMID: 31320610 Free PMC article.
-
Mitophagy regulates mitochondrial network signaling, oxidative stress, and apoptosis during myoblast differentiation.Autophagy. 2019 Sep;15(9):1606-1619. doi: 10.1080/15548627.2019.1591672. Epub 2019 Apr 7. Autophagy. 2019. PMID: 30859901 Free PMC article.
-
Impaired autophagy and APP processing in Alzheimer's disease: The potential role of Beclin 1 interactome.Prog Neurobiol. 2013 Jul-Aug;106-107:33-54. doi: 10.1016/j.pneurobio.2013.06.002. Epub 2013 Jul 1. Prog Neurobiol. 2013. PMID: 23827971 Review.
-
Biological Roles of Alternative Autophagy.Mol Cells. 2018 Jan 31;41(1):50-54. doi: 10.14348/molcells.2018.2215. Epub 2018 Jan 23. Mol Cells. 2018. PMID: 29370693 Free PMC article. Review.
Cited by
-
Integrating miRNA and full-length transcriptome profiling to elucidate the mechanism of muscle growth in Muscovy ducks reveals key roles for miR-301a-3p/ANKRD1.BMC Genomics. 2024 Apr 4;25(1):340. doi: 10.1186/s12864-024-10138-z. BMC Genomics. 2024. PMID: 38575872 Free PMC article.
-
Targeting interleukin-1 for reversing fat browning and muscle wasting in infantile nephropathic cystinosis.J Cachexia Sarcopenia Muscle. 2021 Oct;12(5):1296-1311. doi: 10.1002/jcsm.12744. Epub 2021 Jun 30. J Cachexia Sarcopenia Muscle. 2021. PMID: 34196133 Free PMC article.
-
The role of IL-1 in adipose browning and muscle wasting in CKD-associated cachexia.Sci Rep. 2021 Jul 23;11(1):15141. doi: 10.1038/s41598-021-94565-y. Sci Rep. 2021. PMID: 34302016 Free PMC article.
-
A Leptin Receptor Antagonist Attenuates Adipose Tissue Browning and Muscle Wasting in Infantile Nephropathic Cystinosis-Associated Cachexia.Cells. 2021 Jul 31;10(8):1954. doi: 10.3390/cells10081954. Cells. 2021. PMID: 34440723 Free PMC article.
-
Transcriptome analysis of mRNA and miRNA in the development of LeiZhou goat muscles.Sci Rep. 2024 Apr 29;14(1):9858. doi: 10.1038/s41598-024-60521-9. Sci Rep. 2024. PMID: 38684760 Free PMC article.
References
-
- Lee J.-K., Cho Y.-M., Lee J.-H. Association of Bovine CSRP3 and ACOX1 Genes with Carcass and Meat Quality Traits. Korean J. Agric. Sci. 2010;37:231–238.
-
- Papalouka V., Arvanitis D.A., Vafiadaki E., Mavroidis M., Papadodima S.A., Spiliopoulou C.A., Kremastinos D.T., Kranias E.G., Sanoudou D. Muscle LIM protein interacts with cofilin 2 and regulates F-actin dynamics in cardiac and skeletal muscle. Mol. Cell. Biol. 2009;29:6046–6058. doi: 10.1128/MCB.00654-09. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

