Integrative analysis reveals RNA G-quadruplexes in UTRs are selectively constrained and enriched for functional associations
- PMID: 31988292
- PMCID: PMC6985247
- DOI: 10.1038/s41467-020-14404-y
Integrative analysis reveals RNA G-quadruplexes in UTRs are selectively constrained and enriched for functional associations
Abstract
G-quadruplex (G4) sequences are abundant in untranslated regions (UTRs) of human messenger RNAs, but their functional importance remains unclear. By integrating multiple sources of genetic and genomic data, we show that putative G-quadruplex forming sequences (pG4) in 5' and 3' UTRs are selectively constrained, and enriched for cis-eQTLs and RNA-binding protein (RBP) interactions. Using over 15,000 whole-genome sequences, we find that negative selection acting on central guanines of UTR pG4s is comparable to that of missense variation in protein-coding sequences. At multiple GWAS-implicated SNPs within pG4 UTR sequences, we find robust allelic imbalance in gene expression across diverse tissue contexts in GTEx, suggesting that variants affecting G-quadruplex formation within UTRs may also contribute to phenotypic variation. Our results establish UTR G4s as important cis-regulatory elements and point to a link between disruption of UTR pG4 and disease.
Conflict of interest statement
The authors declare no competing interests.
Figures
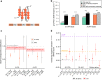




References
MeSH terms
Substances
Grants and funding
- R01 GM128096/GM/NIGMS NIH HHS/United States
- T32 HG000046/HG/NHGRI NIH HHS/United States
- 5T32HG000046-20/U.S. Department of Health & Human Services | NIH | National Human Genome Research Institute (NHGRI)
- GM128096/U.S. Department of Health & Human Services | NIH | National Institute of General Medical Sciences (NIGMS)
LinkOut - more resources
Full Text Sources
Other Literature Sources

