Metabolic Control over mTOR-Dependent Diapause-like State
- PMID: 31991105
- PMCID: PMC7204393
- DOI: 10.1016/j.devcel.2019.12.018
Metabolic Control over mTOR-Dependent Diapause-like State
Abstract
Regulation of embryonic diapause, dormancy that interrupts the tight connection between developmental stage and time, is still poorly understood. Here, we characterize the transcriptional and metabolite profiles of mouse diapause embryos and identify unique gene expression and metabolic signatures with activated lipolysis, glycolysis, and metabolic pathways regulated by AMPK. Lipolysis is increased due to mTORC2 repression, increasing fatty acids to support cell survival. We further show that starvation in pre-implantation ICM-derived mouse ESCs induces a reversible dormant state, transcriptionally mimicking the in vivo diapause stage. During starvation, Lkb1, an upstream kinase of AMPK, represses mTOR, which induces a reversible glycolytic and epigenetically H4K16Ac-negative, diapause-like state. Diapause furthermore activates expression of glutamine transporters SLC38A1/2. We show by genetic and small molecule inhibitors that glutamine transporters are essential for the H4K16Ac-negative, diapause state. These data suggest that mTORC1/2 inhibition, regulated by amino acid levels, is causal for diapause metabolism and epigenetic state.
Keywords: H4K16Ac; LKB1; amino acids; diapause; epigenetics; glutamine transporter; lipolysis; mTOR; metabolism; pluripotent stem cells.
Copyright © 2020 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of Interests The authors declare no competing interests.
Figures



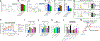

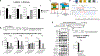

References
-
- Alexa A, Rahnenfuhrer J, and Lengauer T. (2006). Improved scoring of func-€ tional groups from gene expression data by decorrelating GO graph structure. Bioinformatics 22, 1600–1607. - PubMed
-
- Altman BJ, Stine ZE, and Dang CV (2016). From Krebs to clinic: glutamine metabolism to cancer therapy. Nat. Rev. Cancer 16, 749. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

