The evolutionary arms race between transposable elements and piRNAs in Drosophila melanogaster
- PMID: 31992188
- PMCID: PMC6988346
- DOI: 10.1186/s12862-020-1580-3
The evolutionary arms race between transposable elements and piRNAs in Drosophila melanogaster
Abstract
Background: The piwi-interacting RNAs (piRNAs) are small non-coding RNAs that specifically repress transposable elements (TEs) in the germline of Drosophila. Despite our expanding understanding of TE:piRNA interaction, whether there is an evolutionary arms race between TEs and piRNAs was unclear.
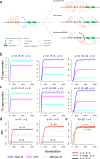
Results: Here, we studied the population genomics of TEs and piRNAs in the worldwide strains of D. melanogaster. By conducting a correlation analysis between TE contents and the abundance of piRNAs from ovaries of representative strains of D. melanogaster, we find positive correlations between TEs and piRNAs in six TE families. Our simulations further highlight that TE activities and the strength of purifying selection against TEs are important factors shaping the interactions between TEs and piRNAs. Our studies also suggest that the de novo generation of piRNAs is an important mechanism to repress the newly invaded TEs.
Conclusions: Our results revealed the existence of an evolutionary arms race between the copy numbers of TEs and the abundance of antisense piRNAs at the population level. Although the interactions between TEs and piRNAs are complex and many factors should be considered to impact their interaction dynamics, our results suggest the emergence, repression specificity and strength of piRNAs on TEs should be considered in studying the landscapes of TE insertions in Drosophila. These results deepen our understanding of the interactions between piRNAs and TEs, and also provide novel insights into the nature of genomic conflicts of other forms.
Keywords: Arms race; Co-evolution; Drosophila melanogaster; Transposable element; piRNA.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures




Similar articles
-
Silencing of Transposable Elements by piRNAs in Drosophila: An Evolutionary Perspective.Genomics Proteomics Bioinformatics. 2017 Jun;15(3):164-176. doi: 10.1016/j.gpb.2017.01.006. Epub 2017 Jun 8. Genomics Proteomics Bioinformatics. 2017. PMID: 28602845 Free PMC article. Review.
-
Population dynamics of PIWI-interacting RNAs (piRNAs) and their targets in Drosophila.Genome Res. 2010 Feb;20(2):212-27. doi: 10.1101/gr.095406.109. Epub 2009 Nov 30. Genome Res. 2010. PMID: 19948818 Free PMC article.
-
Analysis of piRNA-mediated silencing of active TEs in Drosophila melanogaster suggests limits on the evolution of host genome defense.Mol Biol Evol. 2013 Aug;30(8):1816-29. doi: 10.1093/molbev/mst081. Epub 2013 Apr 26. Mol Biol Evol. 2013. PMID: 23625890 Free PMC article.
-
Variation in piRNA and transposable element content in strains of Drosophila melanogaster.Genome Biol Evol. 2014 Sep 29;6(10):2786-98. doi: 10.1093/gbe/evu217. Genome Biol Evol. 2014. PMID: 25267446 Free PMC article.
-
Reexamining the P-Element Invasion of Drosophila melanogaster Through the Lens of piRNA Silencing.Genetics. 2016 Aug;203(4):1513-31. doi: 10.1534/genetics.115.184119. Genetics. 2016. PMID: 27516614 Free PMC article. Review.
Cited by
-
Genomic signatures associated with maintenance of genome stability and venom turnover in two parasitoid wasps.Nat Commun. 2022 Oct 27;13(1):6417. doi: 10.1038/s41467-022-34202-y. Nat Commun. 2022. PMID: 36302851 Free PMC article.
-
Selection on synonymous sites: the unwanted transcript hypothesis.Nat Rev Genet. 2024 Jun;25(6):431-448. doi: 10.1038/s41576-023-00686-7. Epub 2024 Jan 31. Nat Rev Genet. 2024. PMID: 38297070 Review.
-
Evolutionary dynamics between transposable elements and their host genomes: mechanisms of suppression and escape.Curr Opin Genet Dev. 2023 Oct;82:102092. doi: 10.1016/j.gde.2023.102092. Epub 2023 Jul 28. Curr Opin Genet Dev. 2023. PMID: 37517354 Free PMC article. Review.
-
Synergistic epistasis of the deleterious effects of transposable elements.Genetics. 2022 Feb 4;220(2):iyab211. doi: 10.1093/genetics/iyab211. Genetics. 2022. PMID: 34888644 Free PMC article.
-
Co-option of an endogenous retrovirus (LTR7-HERVH) in early human embryogenesis: becoming useful and going unnoticed.Mob DNA. 2025 Jul 5;16(1):27. doi: 10.1186/s13100-025-00361-0. Mob DNA. 2025. PMID: 40618134 Free PMC article. Review.
References
-
- Van Valen L. A new evolutionary law. Evol Theory. 1973;1:1–30.
-
- Neiman M, Fields P. Antagonistic interspecific coevolution. In: Kliman R, editor. Encyclopedia of Evolutionary Biology. Oxford: Academic Press; 2016. pp. 93–100.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

