Double-Stranded RNAs in Plant Protection Against Pathogenic Organisms and Viruses in Agriculture
- PMID: 31993231
- PMCID: PMC6977960
- DOI: 10.32607/20758251-2019-11-4-13-21
Double-Stranded RNAs in Plant Protection Against Pathogenic Organisms and Viruses in Agriculture
Abstract
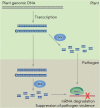
Recent studies have shown that plants are able to express the artificial genes responsible for the synthesis of double-stranded RNAs (dsRNAs) and hairpin double-stranded RNAs (hpRNAs), as well as uptake and process exogenous dsRNAs and hpRNAs to suppress the gene expression of plant pathogenic viruses, fungi, or insects. Both endogenous and exogenous dsRNAs are processed into small interfering RNAs (siRNAs) that can spread locally and systemically through the plant, enter pathogenic microorganisms, and induce RNA interference-mediated pathogen resistance in plants. There are numerous examples of the development of new biotechnological approaches to plant protection using transgenic plants and exogenous dsRNAs. This review summarizes new data on the use of transgenes and exogenous dsRNAs for the suppression of fungal and insect virulence genes, as well as viruses to increase the resistance of plants to these pathogens. We also analyzed the current ideas about the mechanisms of dsRNA processing and transport in plants.
Keywords: RNA interference; double-stranded RNA; exogenous dsRNA; hairpin RNA; plant resistance; regulation of pathogen genes; transgenic plants.
Copyright ® 2019 National Research University Higher School of Economics.
Figures

References
LinkOut - more resources
Full Text Sources
Other Literature Sources
