Diversity and Emerging Roles of Enhancer RNA in Regulation of Gene Expression and Cell Fate
- PMID: 31993419
- PMCID: PMC6971116
- DOI: 10.3389/fcell.2019.00377
Diversity and Emerging Roles of Enhancer RNA in Regulation of Gene Expression and Cell Fate
Abstract
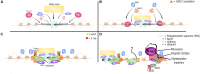
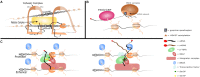
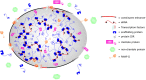
Enhancers are cis-regulatory elements in the genome that cooperate with promoters to control target gene transcription. Unlike promoters, enhancers are not necessarily adjacent to target genes and can exert their functions regardless of enhancer orientations, positions and spatial segregations from target genes. Thus, for a long time, the question as to how enhancers act in a temporal and spatial manner attracted considerable attention. The recent discovery that enhancers are also abundantly transcribed raises interesting questions about the exact roles of enhancer RNA (eRNA) in gene regulation. In this review, we highlight the process of enhancer transcription and the diverse features of eRNA. We review eRNA functions, which include enhancer-promoter looping, chromatin modifying, and transcription regulating. As eRNA are transcribed from active enhancers, they exhibit tissue and lineage specificity, and serve as markers of cell state and function. Finally, we discuss the unique relationship between eRNA and super enhancers in phase separation wherein eRNA may contribute significantly to cell fate decisions.
Keywords: eRNA; enhancer; gene regulation; non-coding RNA; phase separation; super enhancer.
Copyright © 2020 Arnold, Wells and Li.
Figures
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

