Hormonal and gene dynamics in de novo shoot meristem formation during adventitious caulogenesis in cotyledons of Pinus pinea
- PMID: 31993729
- PMCID: PMC7067738
- DOI: 10.1007/s00299-020-02508-0
Hormonal and gene dynamics in de novo shoot meristem formation during adventitious caulogenesis in cotyledons of Pinus pinea
Abstract
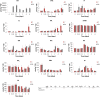
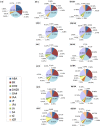
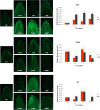
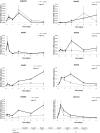
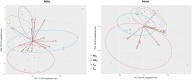
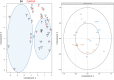
Several members of WOX and KNOX gene families and several plant growth regulators, basically cytokinins and auxins, play a key role during adventitious caulogenesis in the conifer Pinus pinea. Similar to Arabidopsis thaliana, Pinus pinea shoot organogenesis is a multistep process. However, there are key differences between both species, which may alter the underlying physiological and genetic programs. It is unknown if the genic expression models during angiosperm development may be applicable to conifers. In this work, an analysis of the endogenous content of different plant growth regulators and the expression of genes putatively involved in adventitious caulogenesis in P. pinea cotyledons was conducted. A multivariate analysis of both datasets was also realized through partial least squares regression and principal component analysis to obtain an integral vision of the mechanisms involved in caulogenesis in P. pinea. Analyses show that cotyledons cultured in the presence of benzyladenine during long times (2-6 days) cluster separately from the rest of the samples, suggesting that the benzyladenine increase observed during the first hours of culture is sufficient to trigger the caulogenic response through the activation of specific developmental programs. In particular, the most relevant factors involved in this process are the cytokinins trans-zeatin, dihydrozeatin, trans-zeatin riboside and isopentenyl adenosine; the auxin indoleacetic acid; and the genes PpWUS, PpWOX5, PpKN2, PpKN3 and PipiRR1. WUS is functional in pines and has an important role in caulogenesis. Interestingly, WOX5 also seems to participate in the process, although its specific role has not been determined.
Keywords: Conifers; De novo shoot organogenesis; Gene expression; Multivariate analyses; Plant growth regulators.
Conflict of interest statement
The authors declare that they have no conflict of interest. All authors revised and approved the final manuscript.
Figures
References
-
- Aitken-Christie J, Singh AP, Davies H (1988) Multiplication of meristematic tissue: a new tissue culture system for radiata pine. In: Hanover IW, Keathley DE (eds) Genetic manipulation of woody plants. Plenum Press, New York, pp 413–432. 10.1007/978-1-4613-1661-9_26.
-
- Alonso P, Moncaleán P, Fernández B, Rodríguez A, Centeno ML, Ordás RJ. An improved micropropagation protocol for stone pine (Pinus pinea L.) Ann For Sci. 2006;63:879–885.
-
- Alonso P, Cortizo M, Cantón FR, Fernández B, Rodríguez A, Centeno ML, Cánovas FM, Ordás RJ. Identification of genes differentially expressed during adventitious shoot induction in Pinus pinea cotyledons by subtractive hybridization and quantitative PCR. Tree Physiol. 2007;27:1721–1730. - PubMed
-
- Alvarez JM, Cortizo M, Bueno N, Rodríguez A, Ordás RJ. CLAVATA1-LIKE, a leucine-rich-repeat protein receptor kinase gene differentially expressed during adventitious caulogenesis in Pinus pinaster and Pinus pinea. Plant Cell Tiss Org. 2013;112(3):331–342.
-
- Alvarez JM, Bueno N, Cañas RA, Avila C, Cánovas FM, Ordás RJ. Analysis of the WUSCHEL-RELATED HOMEOBOX gene family in Pinus pinaster: new insights into the gene family evolution. Plant Physiol Biochem. 2018;123:304–318. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

