Arginine metabolic control of airway inflammation
- PMID: 31996482
- PMCID: PMC7098726
- DOI: 10.1172/jci.insight.127801
Arginine metabolic control of airway inflammation
Abstract
Inducible nitric oxide synthase (iNOS) and arginase-2 (ARG2) share a common substrate, arginine. Higher expression of iNOS and exhaled NO are linked to airway inflammation in patients. iNOS deletion in animal models suggests that eosinophilic inflammation is regulated by arginine metabolism. Moreover, ARG2 is a regulator of Th2 response, as shown by the development of severe eosinophilic inflammation in ARG2-/- mice. However, potential synergistic roles of iNOS and ARG2 in asthma have not been explored. Here, we hypothesized that arginine metabolic fate via iNOS and ARG2 may govern airway inflammation. In an asthma cohort, ARG2 variant genotypes were associated with arginase activity. ARG2 variants with lower arginase activity, combined with levels of exhaled NO, identified a severe asthma phenotype. Airway inflammation was present in WT, ARG2-/-, iNOS-/-, and ARG2-/-/iNOS-/- mice but was greatest in ARG2-/-. Eosinophilic and neutrophilic infiltration in the ARG2-/- mice was abrogated in ARG2-/-/iNOS-/- animals. Similarly, angiogenic airway remodeling was greatest in ARG2-/- mice. Cytokines driving inflammation and remodeling were highest in lungs of asthmatic ARG2-/- mice and lowest in the iNOS-/-. ARG2 metabolism of arginine suppresses inflammation, while iNOS metabolism promotes airway inflammation, supporting a central role for arginine metabolic control of inflammation.
Keywords: Amino acid metabolism; Asthma; Inflammation; Metabolism; Nitric oxide.
Conflict of interest statement
Figures

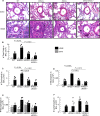



References
-
- Morris SM. Arginine metabolism: boundaries of our knowledge. J Nutr. 2007;137(6 Suppl 2):1602S–1609S. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

