Modeling medulloblastoma in vivo and with human cerebellar organoids
- PMID: 31996670
- PMCID: PMC6989674
- DOI: 10.1038/s41467-019-13989-3
Modeling medulloblastoma in vivo and with human cerebellar organoids
Abstract
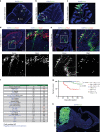
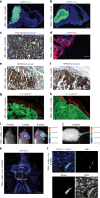
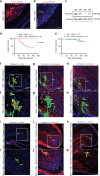
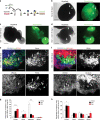
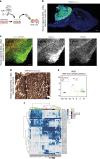
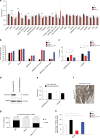
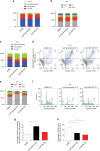
Medulloblastoma (MB) is the most common malignant brain tumor in children and among the subtypes, Group 3 MB has the worst outcome. Here, we perform an in vivo, patient-specific screen leading to the identification of Otx2 and c-MYC as strong Group 3 MB inducers. We validated our findings in human cerebellar organoids where Otx2/c-MYC give rise to MB-like organoids harboring a DNA methylation signature that clusters with human Group 3 tumors. Furthermore, we show that SMARCA4 is able to reduce Otx2/c-MYC tumorigenic activity in vivo and in human cerebellar organoids while SMARCA4 T910M, a mutant form found in human MB patients, inhibits the wild-type protein function. Finally, treatment with Tazemetostat, a EZH2-specific inhibitor, reduces Otx2/c-MYC tumorigenesis in ex vivo culture and human cerebellar organoids. In conclusion, human cerebellar organoids can be efficiently used to understand the role of genes found altered in cancer patients and represent a reliable tool for developing personalized therapies.
Conflict of interest statement
The authors declare no competing interests.
Figures
Similar articles
-
Joint binding of OTX2 and MYC in promotor regions is associated with high gene expression in medulloblastoma.PLoS One. 2011;6(10):e26058. doi: 10.1371/journal.pone.0026058. Epub 2011 Oct 10. PLoS One. 2011. PMID: 22016811 Free PMC article.
-
A group 3 medulloblastoma stem cell program is maintained by OTX2-mediated alternative splicing.Nat Cell Biol. 2024 Aug;26(8):1233-1246. doi: 10.1038/s41556-024-01460-5. Epub 2024 Jul 18. Nat Cell Biol. 2024. PMID: 39025928 Free PMC article.
-
Inactivation of Ezh2 Upregulates Gfi1 and Drives Aggressive Myc-Driven Group 3 Medulloblastoma.Cell Rep. 2017 Mar 21;18(12):2907-2917. doi: 10.1016/j.celrep.2017.02.073. Cell Rep. 2017. PMID: 28329683 Free PMC article.
-
A Proteogenomic Approach to Understanding MYC Function in Metastatic Medulloblastoma Tumors.Int J Mol Sci. 2016 Oct 19;17(10):1744. doi: 10.3390/ijms17101744. Int J Mol Sci. 2016. PMID: 27775567 Free PMC article. Review.
-
Medulloblastoma stem cells: modeling tumor heterogeneity.Cancer Lett. 2013 Sep 10;338(1):23-31. doi: 10.1016/j.canlet.2012.07.010. Epub 2012 Jul 14. Cancer Lett. 2013. PMID: 22796365 Review.
Cited by
-
Exposure of Human Skin Organoids to Low Genotoxic Stress Can Promote Epithelial-to-Mesenchymal Transition in Regenerating Keratinocyte Precursor Cells.Cells. 2020 Aug 18;9(8):1912. doi: 10.3390/cells9081912. Cells. 2020. PMID: 32824646 Free PMC article.
-
Modeling Brain Tumors: A Perspective Overview of in vivo and Organoid Models.Front Mol Neurosci. 2022 May 30;15:818696. doi: 10.3389/fnmol.2022.818696. eCollection 2022. Front Mol Neurosci. 2022. PMID: 35706426 Free PMC article. Review.
-
Organoids: The current status and biomedical applications.MedComm (2020). 2023 May 17;4(3):e274. doi: 10.1002/mco2.274. eCollection 2023 Jun. MedComm (2020). 2023. PMID: 37215622 Free PMC article. Review.
-
Organoids as a new approach for improving pediatric cancer research.Front Oncol. 2024 May 21;14:1414311. doi: 10.3389/fonc.2024.1414311. eCollection 2024. Front Oncol. 2024. PMID: 38835365 Free PMC article. Review.
-
Modeling cancer progression using human pluripotent stem cell-derived cells and organoids.Stem Cell Res. 2020 Dec;49:102063. doi: 10.1016/j.scr.2020.102063. Epub 2020 Oct 27. Stem Cell Res. 2020. PMID: 33137568 Free PMC article. Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

