Flow Sorting Enrichment and Nanopore Sequencing of Chromosome 1 From a Chinese Individual
- PMID: 31998370
- PMCID: PMC6962354
- DOI: 10.3389/fgene.2019.01315
Flow Sorting Enrichment and Nanopore Sequencing of Chromosome 1 From a Chinese Individual
Abstract
Sorting of individual chromosomes by Flow Cytometry (flow-sorting) is an enrichment method to potentially simplify genome assembly by isolating chromosomes from the context of the genome. We have recently developed a workflow to sequence native, unamplified DNA and applied it to the smallest human chromosome, the Y chromosome. Here, we modify improve upon that workflow to increase DNA recovery from chromosome sorting as well as sequencing yield. We apply it to sequence and assemble the largest human chromosome - chromosome 1 - of a Chinese individual using a single Oxford Nanopore MinION flow cell. We generate a selective and highly continuous assembly whose continuity reaches into the order of magnitude of the human reference GRCh38. We then use this assembly to call candidate structural variants against the reference and find 685 putative novel SV candidates. We propose this workflow as a potential solution to assemble structurally complex chromosomes, or the study of very large plant or animal genomes that might challenge traditional assembly strategies.
Keywords: chromosome enrichment; chromosome sequencing; chromosome sorting; flow karyotyping; genome assembly; nanopore sequencing; structural variation.
Copyright © 2020 Kuderna, Solís-Moruno, Batlle-Masó, Julià, Lizano, Anglada, Ramírez, Bote, Tormo, Marquès-Bonet, Fornas and Casals.
Figures
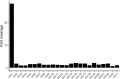
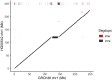

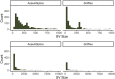
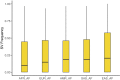
References
LinkOut - more resources
Full Text Sources
Miscellaneous

