Host engulfment pathway controls inflammation in inflammatory bowel disease
- PMID: 32003126
- PMCID: PMC7390677
- DOI: 10.1111/febs.15236
Host engulfment pathway controls inflammation in inflammatory bowel disease
Abstract
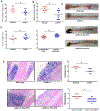
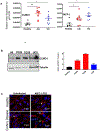
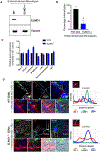
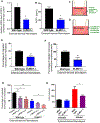
Chronic diseases, including inflammatory bowel disease (IBD) urgently need new biomarkers as a significant proportion of patients, do not respond to current medications. Inflammation is a common factor in these diseases, and microbial sensing in the intestinal tract is critical to initiate the inflammation. We have identified ELMO1 (engulfment and cell motility protein 1) as a microbial sensor in epithelial and phagocytic cells that turns on inflammatory signals. Using a stem cell-based 'gut-in-a-dish' coculture model, we studied the interactions between microbes, epithelium, and monocytes in the context of IBD. To mimic the in vivo cell physiology, enteroid-derived monolayers (EDMs) were generated from the organoids isolated from WT and ELMO1-/- mice and colonic biopsies of IBD patients. The EDMs were infected with the IBD-associated microbes to monitor the inflammatory responses. ELMO1-depleted EDMs displayed a significant reduction in bacterial internalization, a decrease in pro-inflammatory cytokine productions and monocyte recruitment. The expression of ELMO1 is elevated in the colonic epithelium and in the inflammatory infiltrates within the lamina propria of IBD patients where the higher expression is positively correlated with the elevated expression of pro-inflammatory cytokines, MCP-1 and TNF-α. MCP-1 is released from the epithelium and recruits monocytes to the site of inflammation. Once recruited, monocytes require ELMO1 to engulf the bacteria and propagate a robust TNF-α storm. These findings highlight that the dysregulated epithelial ELMO1 → MCP-1 axis can serve as an early biomarker in the diagnostics of IBD and other inflammatory disorders.
Keywords: enteroid; epithelial-immune cell crosstalk; inflammatory bowel disease; microbial sensing and bacterial engulfment; monocyte chemoattractant protein 1.
© 2020 University of California. The FEBS Journal published by John Wiley & Sons Ltd on behalf of Federation of European Biochemical Societies.
Conflict of interest statement
Figures



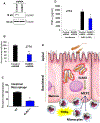
Similar articles
-
Development, validation and implementation of an in vitro model for the study of metabolic and immune function in normal and inflamed human colonic epithelium.Dan Med J. 2015 Jan;62(1):B4973. Dan Med J. 2015. PMID: 25557335 Review.
-
Engulfment and Cell Motility Protein 1 Protects Against DSS-induced Colonic Injury in Mice via Rac1 Activation.J Crohns Colitis. 2019 Jan 1;13(1):100-114. doi: 10.1093/ecco-jcc/jjy133. J Crohns Colitis. 2019. PMID: 30219846
-
The crosstalk between microbial sensors ELMO1 and NOD2 shape intestinal immune responses.Virulence. 2023 Dec;14(1):2171690. doi: 10.1080/21505594.2023.2171690. Virulence. 2023. PMID: 36694274 Free PMC article.
-
Activation of Epithelial Signal Transducer and Activator of Transcription 1 by Interleukin 28 Controls Mucosal Healing in Mice With Colitis and Is Increased in Mucosa of Patients With Inflammatory Bowel Disease.Gastroenterology. 2017 Jul;153(1):123-138.e8. doi: 10.1053/j.gastro.2017.03.015. Epub 2017 Mar 23. Gastroenterology. 2017. PMID: 28342759
-
Dysregulation of Intestinal Epithelial Cell RIPK Pathways Promotes Chronic Inflammation in the IBD Gut.Front Immunol. 2019 May 20;10:1094. doi: 10.3389/fimmu.2019.01094. eCollection 2019. Front Immunol. 2019. PMID: 31164887 Free PMC article. Review.
Cited by
-
Hepatitis E Virus Mediates Renal Injury via the Interaction between the Immune Cells and Renal Epithelium.Vaccines (Basel). 2020 Aug 14;8(3):454. doi: 10.3390/vaccines8030454. Vaccines (Basel). 2020. PMID: 32824088 Free PMC article.
-
eQTL in diseased colon tissue identifies novel target genes associated with IBD.bioRxiv [Preprint]. 2024 Oct 17:2024.10.14.618229. doi: 10.1101/2024.10.14.618229. bioRxiv. 2024. PMID: 39464142 Free PMC article. Preprint.
-
Engulfment and cell motility protein 1 fosters reprogramming of tumor-associated macrophages in colorectal cancer.Cancer Sci. 2023 Feb;114(2):410-422. doi: 10.1111/cas.15628. Epub 2022 Dec 7. Cancer Sci. 2023. PMID: 36310143 Free PMC article.
-
The Use of Gut Organoids: To Study the Physiology and Disease of the Gut Microbiota.J Cell Mol Med. 2025 Feb;29(4):e70330. doi: 10.1111/jcmm.70330. J Cell Mol Med. 2025. PMID: 39968926 Free PMC article. Review.
-
A Novel Strategy to Study the Invasive Capability of Adherent-Invasive Escherichia coli by Using Human Primary Organoid-Derived Epithelial Monolayers.Front Immunol. 2021 Mar 29;12:646906. doi: 10.3389/fimmu.2021.646906. eCollection 2021. Front Immunol. 2021. PMID: 33854511 Free PMC article.
References
-
- Moustafa A, Li W, Anderson EL, Wong EHM, Dulai PS, Sandborn WJ, Biggs W, Yooseph S, Jones MB, Venter JC, Nelson KE, Chang JT, Telenti A & Boland BS (2018) Genetic risk, dysbiosis, and treatment stratification using host genome and gut microbiome in inflammatory bowel disease, Clin Transl Gastroenterol. 9, e132. - PMC - PubMed
-
- Manichanh C, Borruel N, Casellas F & Guarner F (2012) The gut microbiota in IBD, Nat Rev Gastroenterol Hepatol. 9, 599–608. - PubMed
-
- Hugot JP, Chamaillard M, Zouali H, Lesage S, Cezard JP, Belaiche J, Almer S, Tysk C, O'Morain CA, Gassull M, Binder V, Finkel Y, Cortot A, Modigliani R, Laurent-Puig P, Gower-Rousseau C, Macry J, Colombel JF, Sahbatou M & Thomas G (2001) Association of NOD2 leucine-rich repeat variants with susceptibility to Crohn's disease, Nature. 411, 599–603. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

