The full-length structure of Thermus scotoductus OLD defines the ATP hydrolysis properties and catalytic mechanism of Class 1 OLD family nucleases
- PMID: 32009148
- PMCID: PMC7049728
- DOI: 10.1093/nar/gkaa059
The full-length structure of Thermus scotoductus OLD defines the ATP hydrolysis properties and catalytic mechanism of Class 1 OLD family nucleases
Abstract
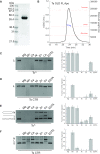
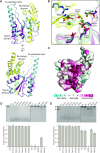
OLD family nucleases contain an N-terminal ATPase domain and a C-terminal Toprim domain. Homologs segregate into two classes based on primary sequence length and the presence/absence of a unique UvrD/PcrA/Rep-like helicase gene immediately downstream in the genome. Although we previously defined the catalytic machinery controlling Class 2 nuclease cleavage, degenerate conservation of the C-termini between classes precludes pinpointing the analogous residues in Class 1 enzymes by sequence alignment alone. Our Class 2 structures also provide no information on ATPase domain architecture and ATP hydrolysis. Here we present the full-length structure of the Class 1 OLD nuclease from Thermus scotoductus (Ts) at 2.20 Å resolution, which reveals a dimerization domain inserted into an N-terminal ABC ATPase fold and a C-terminal Toprim domain. Structural homology with genome maintenance proteins identifies conserved residues responsible for Ts OLD ATPase activity. Ts OLD lacks the C-terminal helical domain present in Class 2 OLD homologs yet preserves the spatial organization of the nuclease active site, arguing that OLD proteins use a conserved catalytic mechanism for DNA cleavage. We also demonstrate that mutants perturbing ATP hydrolysis or DNA cleavage in vitro impair P2 OLD-mediated killing of recBC-Escherichia coli hosts, indicating that both the ATPase and nuclease activities are required for OLD function in vivo.
© The Author(s) 2020. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures




Similar articles
-
OLD family nuclease function across diverse anti-phage defense systems.Front Microbiol. 2023 Sep 28;14:1268820. doi: 10.3389/fmicb.2023.1268820. eCollection 2023. Front Microbiol. 2023. PMID: 37840731 Free PMC article. Review.
-
Structural characterization of Class 2 OLD family nucleases supports a two-metal catalysis mechanism for cleavage.Nucleic Acids Res. 2019 Sep 26;47(17):9448-9463. doi: 10.1093/nar/gkz703. Nucleic Acids Res. 2019. PMID: 31400118 Free PMC article.
-
Mechanism of DnaB helicase of Escherichia coli: structural domains involved in ATP hydrolysis, DNA binding, and oligomerization.Biochemistry. 1999 Aug 24;38(34):10919-28. doi: 10.1021/bi990048t. Biochemistry. 1999. PMID: 10460147
-
Cooperation of the N-terminal Helicase and C-terminal endonuclease activities of Archaeal Hef protein in processing stalled replication forks.J Biol Chem. 2004 Dec 17;279(51):53175-85. doi: 10.1074/jbc.M409243200. Epub 2004 Oct 12. J Biol Chem. 2004. PMID: 15485882
-
Research progress of the SLFN family in malignant tumors.Front Oncol. 2024 Nov 4;14:1468484. doi: 10.3389/fonc.2024.1468484. eCollection 2024. Front Oncol. 2024. PMID: 39558948 Free PMC article. Review.
Cited by
-
Structural and biochemical insights into the mechanism of the Gabija bacterial immunity system.Nat Commun. 2024 Jan 29;15(1):836. doi: 10.1038/s41467-024-45173-7. Nat Commun. 2024. PMID: 38282040 Free PMC article.
-
A Class 1 OLD family nuclease encoded by Vibrio cholerae is countered by a vibriophage-encoded direct inhibitor.bioRxiv [Preprint]. 2025 Jan 6:2025.01.06.631583. doi: 10.1101/2025.01.06.631583. bioRxiv. 2025. PMID: 39829814 Free PMC article. Preprint.
-
Comprehensive classification of ABC ATPases and their functional radiation in nucleoprotein dynamics and biological conflict systems.Nucleic Acids Res. 2020 Oct 9;48(18):10045-10075. doi: 10.1093/nar/gkaa726. Nucleic Acids Res. 2020. PMID: 32894288 Free PMC article. Review.
-
A Vibrio cholerae anti-phage system depletes nicotinamide adenine dinucleotide to restrict virulent bacteriophages.mBio. 2024 Nov 13;15(11):e0245724. doi: 10.1128/mbio.02457-24. Epub 2024 Oct 8. mBio. 2024. PMID: 39377576 Free PMC article.
-
OLD family nuclease function across diverse anti-phage defense systems.Front Microbiol. 2023 Sep 28;14:1268820. doi: 10.3389/fmicb.2023.1268820. eCollection 2023. Front Microbiol. 2023. PMID: 37840731 Free PMC article. Review.
References
-
- Koonin E.V., Gorbalenya A.E.. The superfamily of UvrA-related ATPases includes three more subunits of putative ATP-dependent nucleases. Protein Seq Data Anal. 1992; 5:43–45. - PubMed
-
- Sironi G. Mutants of Escherichia coli unable to be lysogenized by the temperate bacteriophage P2. Virology. 1969; 37:163–176. - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources

