Resequencing 200 Flax Cultivated Accessions Identifies Candidate Genes Related to Seed Size and Weight and Reveals Signatures of Artificial Selection
- PMID: 32010166
- PMCID: PMC6976528
- DOI: 10.3389/fpls.2019.01682
Resequencing 200 Flax Cultivated Accessions Identifies Candidate Genes Related to Seed Size and Weight and Reveals Signatures of Artificial Selection
Abstract
Seed size and weight are key traits determining crop yield, which often undergo strongly artificial selection during crop domestication. Although seed sizes differ significantly between oil flax and fiber flax, the genetic basis of morphological differences and artificial selection characteristics in seed size remains largely unclear. Here we re-sequenced 200 flax cultivated accessions to generate a genome variation map based on chromosome assembly reference genomes. We provide evidence that oil flax group is the ancestor of cultivated flax, and the oil-fiber dual purpose group (OF) is the evolutionary intermediate transition state between oil and fiber flax. Genome-wide association studies (GWAS) were combined with LD Heatmap to identify candidate regions related to seed size and weight, then candidate genes were screened based on detailed functional annotations and estimation of nucleotide polymorphism effects. Using this strategy, we obtained 13 candidate genes related to seed size and weight. Selective sweeps analysis indicates human-involved selection of small seeds during the oil to fiber flax transition. Our study shows the existence of elite alleles for seed size and weight in flax germplasm and provides molecular insights into approaches for further improvement.
Keywords: domestication; flax (Linum usitatissimum L.); genetic diversity; genome-wide association studies; seed size; selective sweeps.
Copyright © 2020 Guo, Jiang, Yan, Yang, Ye, Wang, Yan, Chen, Gao, Duan, Liu and Xie.
Figures



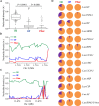
References
-
- Austin D. F. (2012). Domestication of plants in the old world. the origin and spread of domesticated plants in South-west Asia, Europe, and the Mediterranean Basin. Fourth Edition by Daniel Zohary; Maria Hopf; Ehud Weiss. Econ. Bot. 66, 420–421. 10.2307/23325662 - DOI
LinkOut - more resources
Full Text Sources
Research Materials

