Targeting dePARylation for cancer therapy
- PMID: 32010441
- PMCID: PMC6988220
- DOI: 10.1186/s13578-020-0375-y
Targeting dePARylation for cancer therapy
Abstract
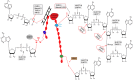
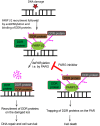
Poly(ADP-ribosyl)ation (PARylation) mediated by poly ADP-ribose polymerases (PARPs) plays a key role in DNA damage repair. Suppression of PARylation by PARP inhibitors impairs DNA damage repair and induces apoptosis of tumor cells with repair defects. Thus, PARP inhibitors have been approved by the US FDA for various types of cancer treatment. However, recent studies suggest that dePARylation also plays a key role in DNA damage repair. Instead of antagonizing PARylation, dePARylation acts as a downstream step of PARylation in DNA damage repair. Moreover, several types of dePARylation inhibitors have been developed and examined in the preclinical studies for cancer treatment. In this review, we will discuss the recent progress on the role of dePARylation in DNA damage repair and cancer suppression. We expect that targeting dePARylation could be a promising approach for cancer chemotherapy in the future.
Keywords: ADP-ribosylation; Cancer therapy; DNA damage response; PARG; dePARylation.
© The Author(s) 2020.
Conflict of interest statement
Competing interestsThe authors declare that they have no competing interests.
Figures
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources

