Genomic evolution of Neisseria gonorrhoeae since the preantibiotic era (1928-2013): antimicrobial use/misuse selects for resistance and drives evolution
- PMID: 32013864
- PMCID: PMC6998845
- DOI: 10.1186/s12864-020-6511-6
Genomic evolution of Neisseria gonorrhoeae since the preantibiotic era (1928-2013): antimicrobial use/misuse selects for resistance and drives evolution
Abstract
Background: Multidrug-resistant Neisseria gonorrhoeae strains are prevalent, threatening gonorrhoea treatment globally, and understanding of emergence, evolution, and spread of antimicrobial resistance (AMR) in gonococci remains limited. We describe the genomic evolution of gonococci and their AMR, related to the introduction of antimicrobial therapies, examining isolates from 1928 (preantibiotic era) to 2013 in Denmark. This is, to our knowledge, the oldest gonococcal collection globally.
Methods: Lyophilised isolates were revived and examined using Etest (18 antimicrobials) and whole-genome sequencing (WGS). Quality-assured genome sequences were obtained for 191 viable and 40 non-viable isolates and analysed with multiple phylogenomic approaches.
Results: Gonococcal AMR, including an accumulation of multiple AMR determinants, started to emerge particularly in the 1950s-1970s. By the twenty-first century, resistance to most antimicrobials was common. Despite that some AMR determinants affect many physiological functions and fitness, AMR determinants were mainly selected by the use/misuse of gonorrhoea therapeutic antimicrobials. Most AMR developed in strains belonging to one multidrug-resistant (MDR) clade with close to three times higher genomic mutation rate. Modern N. gonorrhoeae was inferred to have emerged in the late-1500s and its genome became increasingly conserved over time.
Conclusions: WGS of gonococci from 1928 to 2013 showed that no AMR determinants, except penB, were in detectable frequency before the introduction of gonorrhoea therapeutic antimicrobials. The modern gonococcus is substantially younger than previously hypothesized and has been evolving into a more clonal species, driven by the use/misuse of antimicrobials. The MDR gonococcal clade should be further investigated for early detection of strains with predispositions to develop and maintain MDR and for initiation of public health interventions.
Keywords: Antimicrobial resistance; Evolution; Genomic epidemiology; Neisseria gonorrhoeae; Temporal analysis; Whole-genome sequencing.
Conflict of interest statement
All authors declare that they have no competing interests.
Figures


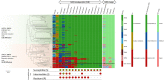



References
-
- Report on global sexually transmitted infection surveillance 2018. World Health Organization (WHO). https://www.who.int/reproductivehealth/publications/stis-surveillance-20.... Accessed 14 Sept 2019.
-
- Unemo Magnus, Lahra Monica M., Cole Michelle, Galarza Patricia, Ndowa Francis, Martin Irene, Dillon Jo-Anne R., Ramon-Pardo Pilar, Bolan Gail, Wi Teodora. World Health Organization Global Gonococcal Antimicrobial Surveillance Program (WHO GASP): review of new data and evidence to inform international collaborative actions and research efforts. Sexual Health. 2019;16(5):412. doi: 10.1071/SH19023. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

