Genetic diversity and phylogenetic structure of four Tibeto-Burman-speaking populations in Tibetan-Yi corridor revealed by insertion/deletion polymorphisms
- PMID: 32017463
- PMCID: PMC7196475
- DOI: 10.1002/mgg3.1140
Genetic diversity and phylogenetic structure of four Tibeto-Burman-speaking populations in Tibetan-Yi corridor revealed by insertion/deletion polymorphisms
Retraction in
-
Retraction: Genetic diversity and phylogenetic structure of four Tibeto-Burman-speaking populations in Tibetan-Yi corridor revealed by insertion/deletion polymorphisms.Mol Genet Genomic Med. 2024 Feb;12(2):e2371. doi: 10.1002/mgg3.2371. Mol Genet Genomic Med. 2024. PMID: 38345203 Free PMC article.
Abstract
Background: Insertion/deletion polymorphisms (InDels), combined with all the desirable features of both short tandem repeat and single nucleotide polymorphism, have been used in archaeological and anthropological research, population genetics and forensic application.
Methods: Thirty InDels in 530 individuals residing in the Tibetan-Yi corridor (142 Dujiangyan Tibetans, 164 Muli Tibetans, 187 Xichang Yis, and 37 Yanyuan Mosuos) were genotyped using the Investigator DIPplex. Forensic parameters and allele frequency spectrum were calculated. Genetic relationships between the investigated populations and worldwide and nationwide populations were assessed based on both the allele frequency distribution and genotype data.
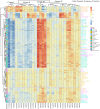
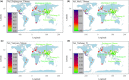
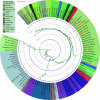
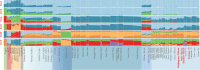
Results: The combined powers of exclusion were 0.9807 (Dujiangyan Tibetan), 0.9880 (Muli Tibetan), 0.9852 (Xichang Yi) and 0.9892 (Yanyuan Mosuo). The combined powers of discrimination were 0.999999999983 (Dujiangyan Tibetan), 0.999999999942 (Muli Tibetan), 0.999999999982 (Xichang Yi) and 0.999999999962 (Yanyuan Mosuo), respectively. The comprehensive population comparisons among worldwide and nationwide populations uniformly illustrated that the investigated populations have a genetically closer relationship with Tibeto-Burman-speaking populations and geographically adjacent populations.
Conclusion: These 30 loci can be regarded as an efficient genetic tool in forensic individual identification and as a supplementary tool in paternity testing in Dujiangyan Tibetan, Muli Tibetan, Xichang Yi, and Yanyuan Mosuo. The genetic proximity between the four populations in the Tibetan-Yi corridor and other populations is strongly correlated with the linguistic origin and geographical distance.
Keywords: InDels; Tibetan-Yi corridor; forensic genetics; phylogenetic reconstruction; population genetics.
© 2020 The Authors. Molecular Genetics & Genomic Medicine published by Wiley Periodicals, Inc.
Conflict of interest statement
The authors have declared that they have no conflict of interest.
Figures


Similar articles
-
Revisiting the genetic background and phylogenetic structure of five Sino-Tibetan-speaking populations: insights from autosomal InDels.Mol Genet Genomics. 2020 Jul;295(4):969-979. doi: 10.1007/s00438-020-01673-x. Epub 2020 Apr 11. Mol Genet Genomics. 2020. PMID: 32279092
-
Population genetics and forensic efficiency of 30 InDel markers in four Chinese ethnic groups residing in Sichuan.Forensic Sci Res. 2020 Apr 21;7(3):498-502. doi: 10.1080/20961790.2020.1737470. eCollection 2022. Forensic Sci Res. 2020. PMID: 36353334 Free PMC article.
-
Genetic diversity, structure and forensic characteristics of Hmong-Mien-speaking Miao revealed by autosomal insertion/deletion markers.Mol Genet Genomics. 2019 Dec;294(6):1487-1498. doi: 10.1007/s00438-019-01591-7. Epub 2019 Jul 16. Mol Genet Genomics. 2019. PMID: 31312894
-
Population genetics, diversity and forensic characteristics of Tai-Kadai-speaking Bouyei revealed by insertion/deletions markers.Mol Genet Genomics. 2019 Oct;294(5):1343-1357. doi: 10.1007/s00438-019-01584-6. Epub 2019 Jun 13. Mol Genet Genomics. 2019. PMID: 31197471
-
Forensic parameters of the Investigator DIPplex kit (Qiagen) in six Mexican populations.Int J Legal Med. 2016 May;130(3):683-5. doi: 10.1007/s00414-015-1242-y. Epub 2015 Aug 2. Int J Legal Med. 2016. PMID: 26233613 Review.
Cited by
-
Genetic variation and phylogenetic analysis of 23 STR in Chinese Han population from Hainan, Southern China.Medicine (Baltimore). 2024 May 31;103(22):e38428. doi: 10.1097/MD.0000000000038428. Medicine (Baltimore). 2024. PMID: 39259071 Free PMC article.
-
Genomic formation of Tibeto-Burman speaking populations in Guizhou, Southwest China.BMC Genomics. 2023 Nov 7;24(1):672. doi: 10.1186/s12864-023-09767-7. BMC Genomics. 2023. PMID: 37936086 Free PMC article.
References
-
- Bouwman, A. S. , Kennedy, S. L. , Muller, R. , Stephens, R. H. , Holst, M. , Caffell, A. C. , … Brown, T. A. (2012). Genotype of a historic strain of Mycobacterium tuberculosis. Proceedings of the National Academy of Sciences of the United States of America, 109, 18511–18516. 10.1073/pnas.1209444109 - DOI - PMC - PubMed
-
- Cummings, M. P. (2004). PHYLIP (phylogeny inference package). New York, NY: John Wiley & Sons Inc.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

