DNA topoisomerase IIIβ promotes cyst generation by inducing cyst wall protein gene expression in Giardia lamblia
- PMID: 32019477
- PMCID: PMC7058931
- DOI: 10.1098/rsob.190228
DNA topoisomerase IIIβ promotes cyst generation by inducing cyst wall protein gene expression in Giardia lamblia
Abstract
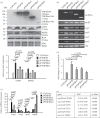
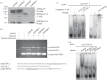
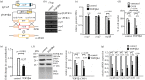
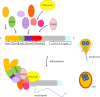
Giardia lamblia causes waterborne diarrhoea by transmission of infective cysts. Three cyst wall proteins are highly expressed in a concerted manner during encystation of trophozoites into cysts. However, their gene regulatory mechanism is still largely unknown. DNA topoisomerases control topological homeostasis of genomic DNA during replication, transcription and chromosome segregation. They are involved in a variety of cellular processes including cell cycle, cell proliferation and differentiation, so they may be valuable drug targets. Giardia lamblia possesses a type IA DNA topoisomerase (TOP3β) with similarity to the mammalian topoisomerase IIIβ. We found that TOP3β was upregulated during encystation and it possessed DNA-binding and cleavage activity. TOP3β can bind to the cwp promoters in vivo using norfloxacin-mediated topoisomerase immunoprecipitation assays. We also found TOP3β can interact with MYB2, a transcription factor involved in the coordinate expression of cwp1-3 genes during encystation. Interestingly, overexpression of TOP3β increased expression of cwp1-3 and myb2 genes and cyst formation. Microarray analysis confirmed upregulation of cwp1-3 and myb2 genes by TOP3β. Mutation of the catalytically important Tyr residue, deletion of C-terminal zinc ribbon domain or further deletion of partial catalytic core domain reduced the levels of cleavage activity, cwp1-3 and myb2 gene expression, and cyst formation. Interestingly, some of these mutant proteins were mis-localized to cytoplasm. Using a CRISPR/Cas9 system for targeted disruption of top3β gene, we found a significant decrease in cwp1-3 and myb2 gene expression and cyst number. Our results suggest that TOP3β may be functionally conserved, and involved in inducing Giardia cyst formation.
Keywords: DNA-binding protein; Giardia; cyst; differentiation; topoisomerase IIIβ; transcription.
Conflict of interest statement
We declare we have no competing interests.
Figures





Similar articles
-
A Novel Multiprotein Bridging Factor 1-Like Protein Induces Cyst Wall Protein Gene Expression and Cyst Differentiation in Giardia lamblia.Int J Mol Sci. 2021 Jan 29;22(3):1370. doi: 10.3390/ijms22031370. Int J Mol Sci. 2021. PMID: 33573049 Free PMC article.
-
A Novel Spo11 Homologue Functions as a Positive Regulator in Cyst Differentiation in Giardia lamblia.Int J Mol Sci. 2021 Nov 2;22(21):11902. doi: 10.3390/ijms222111902. Int J Mol Sci. 2021. PMID: 34769330 Free PMC article.
-
DNA topoisomerase II is involved in regulation of cyst wall protein genes and differentiation in Giardia lamblia.PLoS Negl Trop Dis. 2013 May 16;7(5):e2218. doi: 10.1371/journal.pntd.0002218. Print 2013. PLoS Negl Trop Dis. 2013. PMID: 23696909 Free PMC article.
-
Mechanisms of adaptation in the intestinal parasite Giardia lamblia.Essays Biochem. 2011;51:177-91. doi: 10.1042/bse0510177. Essays Biochem. 2011. PMID: 22023449 Review.
-
Type IA topoisomerases can be "magicians" for both DNA and RNA in all domains of life.RNA Biol. 2017 Jul 3;14(7):854-864. doi: 10.1080/15476286.2017.1330741. Epub 2017 May 23. RNA Biol. 2017. PMID: 28534707 Free PMC article. Review.
Cited by
-
A Novel Multiprotein Bridging Factor 1-Like Protein Induces Cyst Wall Protein Gene Expression and Cyst Differentiation in Giardia lamblia.Int J Mol Sci. 2021 Jan 29;22(3):1370. doi: 10.3390/ijms22031370. Int J Mol Sci. 2021. PMID: 33573049 Free PMC article.
-
Insights into the DNA and RNA Interactions of Human Topoisomerase III Beta Using Molecular Dynamics Simulations.J Chem Inf Model. 2024 Aug 12;64(15):6062-6071. doi: 10.1021/acs.jcim.4c00472. Epub 2024 Jul 18. J Chem Inf Model. 2024. PMID: 39024468 Free PMC article.
-
Target acquired: transcriptional regulators as drug targets for protozoan parasites.Int J Parasitol. 2021 Jul;51(8):599-611. doi: 10.1016/j.ijpara.2020.12.007. Epub 2021 Mar 13. Int J Parasitol. 2021. PMID: 33722681 Free PMC article. Review.
-
Evaluation of the Antiparasitic, Antihepatotoxicity, and Antioxidant Efficacy of Quercetin and Chitosan, Either Alone or in Combination, against Infection Induced by Giardia lamblia in Male Rats.Life (Basel). 2023 Dec 10;13(12):2316. doi: 10.3390/life13122316. Life (Basel). 2023. PMID: 38137916 Free PMC article.
-
A Novel Spo11 Homologue Functions as a Positive Regulator in Cyst Differentiation in Giardia lamblia.Int J Mol Sci. 2021 Nov 2;22(21):11902. doi: 10.3390/ijms222111902. Int J Mol Sci. 2021. PMID: 34769330 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

