From methylglyoxal to pyruvate: a genome-wide study for the identification of glyoxalases and D-lactate dehydrogenases in Sorghum bicolor
- PMID: 32041545
- PMCID: PMC7011430
- DOI: 10.1186/s12864-020-6547-7
From methylglyoxal to pyruvate: a genome-wide study for the identification of glyoxalases and D-lactate dehydrogenases in Sorghum bicolor
Abstract
Background: The glyoxalase pathway is evolutionarily conserved and involved in the glutathione-dependent detoxification of methylglyoxal (MG), a cytotoxic by-product of glycolysis. It acts via two metallo-enzymes, glyoxalase I (GLYI) and glyoxalase II (GLYII), to convert MG into D-lactate, which is further metabolized to pyruvate by D-lactate dehydrogenases (D-LDH). Since D-lactate formation occurs solely by the action of glyoxalase enzymes, its metabolism may be considered as the ultimate step of MG detoxification. By maintaining steady state levels of MG and other reactive dicarbonyl compounds, the glyoxalase pathway serves as an important line of defence against glycation and oxidative stress in living organisms. Therefore, considering the general role of glyoxalases in stress adaptation and the ability of Sorghum bicolor to withstand prolonged drought, the sorghum glyoxalase pathway warrants an in-depth investigation with regard to the presence, regulation and distribution of glyoxalase and D-LDH genes.
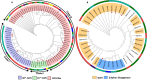
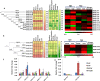
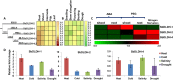
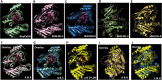
Result: Through this study, we have identified 15 GLYI and 6 GLYII genes in sorghum. In addition, 4 D-LDH genes were also identified, forming the first ever report on genome-wide identification of any plant D-LDH family. Our in silico analysis indicates homology of putatively active SbGLYI, SbGLYII and SbDLDH proteins to several functionally characterised glyoxalases and D-LDHs from Arabidopsis and rice. Further, these three gene families exhibit development and tissue-specific variations in their expression patterns. Importantly, we could predict the distribution of putatively active SbGLYI, SbGLYII and SbDLDH proteins in at least four different sub-cellular compartments namely, cytoplasm, chloroplast, nucleus and mitochondria. Most of the members of the sorghum glyoxalase and D-LDH gene families are indeed found to be highly stress responsive.
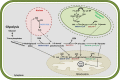
Conclusion: This study emphasizes the role of glyoxalases as well as that of D-LDH in the complete detoxification of MG in sorghum. In particular, we propose that D-LDH which metabolizes the specific end product of glyoxalases pathway is essential for complete MG detoxification. By proposing a cellular model for detoxification of MG via glyoxalase pathway in sorghum, we suggest that different sub-cellular organelles are actively involved in MG metabolism in plants.
Keywords: D-lactate dehydrogenase; Genome-wide analysis; Glyoxalase; Sorghum; Stress.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
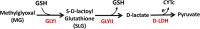



Similar articles
-
Genomic identification, characterization, and stress-induced expression profiling of glyoxalase and D-lactate dehydrogenase gene families in Capsicum annuum.BMC Plant Biol. 2024 Oct 21;24(1):990. doi: 10.1186/s12870-024-05612-5. BMC Plant Biol. 2024. PMID: 39428463 Free PMC article.
-
Pyramiding D-lactate dehydrogenase with the glyoxalase pathway enhances abiotic stress tolerance in plants.Plant Physiol Biochem. 2024 Feb;207:108391. doi: 10.1016/j.plaphy.2024.108391. Epub 2024 Jan 26. Plant Physiol Biochem. 2024. PMID: 38309183
-
Genome-wide analysis and expression profiling of glyoxalase gene families in soybean (Glycine max) indicate their development and abiotic stress specific response.BMC Plant Biol. 2016 Apr 16;16:87. doi: 10.1186/s12870-016-0773-9. BMC Plant Biol. 2016. PMID: 27083416 Free PMC article.
-
Characteristic Variations and Similarities in Biochemical, Molecular, and Functional Properties of Glyoxalases across Prokaryotes and Eukaryotes.Int J Mol Sci. 2017 Mar 30;18(4):250. doi: 10.3390/ijms18040250. Int J Mol Sci. 2017. PMID: 28358304 Free PMC article. Review.
-
An overview on the role of methylglyoxal and glyoxalases in plants.Drug Metabol Drug Interact. 2008;23(1-2):51-68. doi: 10.1515/dmdi.2008.23.1-2.51. Drug Metabol Drug Interact. 2008. PMID: 18533364 Review.
Cited by
-
Physiological and Differential Proteomic Analysis at Seedling Stage by Induction of Heavy-Ion Beam Radiation in Wheat Seeds.Front Genet. 2022 Jul 19;13:942806. doi: 10.3389/fgene.2022.942806. eCollection 2022. Front Genet. 2022. PMID: 35928451 Free PMC article.
-
Glutathione Metabolism in Plants under Stress: Beyond Reactive Oxygen Species Detoxification.Metabolites. 2021 Sep 19;11(9):641. doi: 10.3390/metabo11090641. Metabolites. 2021. PMID: 34564457 Free PMC article. Review.
-
Genomic identification, characterization, and stress-induced expression profiling of glyoxalase and D-lactate dehydrogenase gene families in Capsicum annuum.BMC Plant Biol. 2024 Oct 21;24(1):990. doi: 10.1186/s12870-024-05612-5. BMC Plant Biol. 2024. PMID: 39428463 Free PMC article.
-
Aldehyde dehydrogenase superfamily in sorghum: genome-wide identification, evolution, and transcript profiling during development stages and stress conditions.BMC Plant Biol. 2022 Jul 4;22(1):316. doi: 10.1186/s12870-022-03708-4. BMC Plant Biol. 2022. PMID: 35786175 Free PMC article.
-
Lactate formation from fructose or C1 compounds in the acetogen Acetobacterium woodii by metabolic engineering.Appl Microbiol Biotechnol. 2023 Sep;107(17):5491-5502. doi: 10.1007/s00253-023-12637-7. Epub 2023 Jul 7. Appl Microbiol Biotechnol. 2023. PMID: 37417977 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

