A Circular RNA from the MDM2 Locus Controls Cell Cycle Progression by Suppressing p53 Levels
- PMID: 32041821
- PMCID: PMC7156218
- DOI: 10.1128/MCB.00473-19
A Circular RNA from the MDM2 Locus Controls Cell Cycle Progression by Suppressing p53 Levels
Abstract
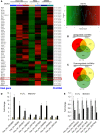
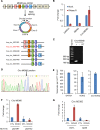
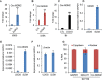
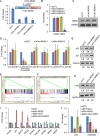
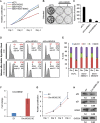
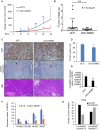
Circular RNAs (circRNAs) are a class of noncoding RNAs produced by a noncanonical form of alternative splicing called back-splicing. To investigate a potential role of circRNAs in the p53 pathway, we analyzed RNA sequencing (RNA-seq) data from colorectal cancer cell lines (HCT116, RKO, and SW48) that were untreated or treated with a DNA-damaging agent. Surprisingly, unlike the strong p53-dependent induction of hundreds of p53-induced mRNAs upon DNA damage, only a few circRNAs were upregulated from p53-induced genes. circ-MDM2, an annotated circRNA from the MDM2 locus, was one of the handful of circRNAs that originated from a p53-induced gene. Given the central role of MDM2 in suppressing p53 protein levels and p53 activity, we investigated the function of circ-MDM2 Knocking down circ-MDM2 with small interfering RNAs (siRNAs) that targeted circ-MDM2 did not alter MDM2 mRNA or MDM2 protein levels but resulted in increased basal p53 levels and growth defects in vitro and in vivo Consistent with these results, transcriptome profiling showed increased expression of several direct p53 targets, reduced retinoblastoma protein (Rb) phosphorylation, and defects in G1-S progression upon silencing circ-MDM2 Our results on the initial characterization of circ-MDM2 identify a new player from the MDM2 locus that suppresses p53 levels and cell cycle progression.
Keywords: CRC; DNA damage; MDM2; cell cycle; circRNA; circular RNA; p53.
Copyright © 2020 American Society for Microbiology.
Figures
Similar articles
-
Circular RNA CDR1as disrupts the p53/MDM2 complex to inhibit Gliomagenesis.Mol Cancer. 2020 Sep 7;19(1):138. doi: 10.1186/s12943-020-01253-y. Mol Cancer. 2020. PMID: 32894144 Free PMC article.
-
Expression profiles of circular RNAs in human colorectal cancer based on RNA deep sequencing.J Clin Lab Anal. 2019 Sep;33(7):e22952. doi: 10.1002/jcla.22952. Epub 2019 Jun 6. J Clin Lab Anal. 2019. PMID: 31169949 Free PMC article.
-
RNA sequencing reveals the expression profiles of circRNA and indicates that circDDX17 acts as a tumor suppressor in colorectal cancer.J Exp Clin Cancer Res. 2018 Dec 27;37(1):325. doi: 10.1186/s13046-018-1006-x. J Exp Clin Cancer Res. 2018. PMID: 30591054 Free PMC article.
-
[Any way you splice it: Mdm2 at the crossroads of tumor surveillance].Ai Zheng. 2008 Sep;27(9):993-7. Ai Zheng. 2008. PMID: 18799043 Review. Chinese.
-
The potential role of circular RNAs in regulating p53 in different types of cancers.Pathol Res Pract. 2024 Sep;261:155488. doi: 10.1016/j.prp.2024.155488. Epub 2024 Jul 25. Pathol Res Pract. 2024. PMID: 39088876 Review.
Cited by
-
The Role of Circular RNAs in DNA Damage Response and Repair.Cancers (Basel). 2021 Oct 26;13(21):5352. doi: 10.3390/cancers13215352. Cancers (Basel). 2021. PMID: 34771517 Free PMC article. Review.
-
Nanoparticles-Delivered Circular RNA Strategy as a Novel Antitumor Approach.Int J Mol Sci. 2024 Aug 16;25(16):8934. doi: 10.3390/ijms25168934. Int J Mol Sci. 2024. PMID: 39201617 Free PMC article. Review.
-
Non-coding RNAs participate in interactions between senescence and gastrointestinal cancers.Front Genet. 2025 Jan 3;15:1461404. doi: 10.3389/fgene.2024.1461404. eCollection 2024. Front Genet. 2025. PMID: 39831201 Free PMC article.
-
The multifaceted roles of circular RNAs in cancer hallmarks: From mechanisms to clinical implications.Mol Ther Nucleic Acids. 2024 Jul 20;35(3):102286. doi: 10.1016/j.omtn.2024.102286. eCollection 2024 Sep 10. Mol Ther Nucleic Acids. 2024. PMID: 39188305 Free PMC article. Review.
-
Circular RNAs regulate cancer-related signaling pathways and serve as potential diagnostic biomarkers for human cancers.Cancer Cell Int. 2021 Jun 23;21(1):317. doi: 10.1186/s12935-021-02017-4. Cancer Cell Int. 2021. PMID: 34162394 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous
