Multistep peripherin-2/rds self-assembly drives membrane curvature for outer segment disk architecture and photoreceptor viability
- PMID: 32041874
- PMCID: PMC7049111
- DOI: 10.1073/pnas.1912513117
Multistep peripherin-2/rds self-assembly drives membrane curvature for outer segment disk architecture and photoreceptor viability
Abstract
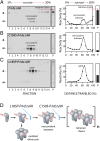
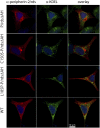
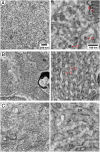
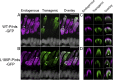
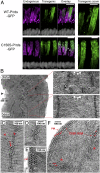
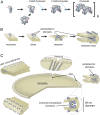
Rod and cone photoreceptor outer segment (OS) structural integrity is essential for normal vision; disruptions contribute to a broad variety of retinal ciliopathies. OSs possess many hundreds of stacked membranous disks, which capture photons and scaffold the phototransduction cascade. Although the molecular basis of OS structure remains unresolved, recent studies suggest that the photoreceptor-specific tetraspanin, peripherin-2/rds (P/rds), may contribute to the highly curved rim domains at disk edges. Here, we demonstrate that tetrameric P/rds self-assembly is required for generating high-curvature membranes in cellulo, implicating the noncovalent tetramer as a minimal unit of function. P/rds activity was promoted by disulfide-mediated tetramer polymerization, which transformed localized regions of curvature into high-curvature tubules of extended lengths. Transmission electron microscopy visualization of P/rds purified from OS membranes revealed disulfide-linked tetramer chains up to 100 nm long, suggesting that chains maintain membrane curvature continuity over extended distances. We tested this idea in Xenopus laevis photoreceptors, and found that transgenic expression of nonchain-forming P/rds generated abundant high-curvature OS membranes, which were improperly but specifically organized as ectopic incisures and disk rims. These striking phenotypes demonstrate the importance of P/rds tetramer chain formation for the continuity of rim formation during disk morphogenesis. Overall, this study advances understanding of the normal structure and function of P/rds for OS architecture and biogenesis, and clarifies how pathogenic loss-of-function mutations in P/rds cause photoreceptor structural defects to trigger progressive retinal degenerations. It also introduces the possibility that other tetraspanins may generate or sense membrane curvature in support of diverse biological functions.
Keywords: cilium; digenic retinitis pigmentosa; membrane curvature; photoreceptor outer segment; tetraspanin.
Conflict of interest statement
The authors declare no competing interest.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

