Point Mutations in Retargeted gD Eliminate the Sensitivity of EGFR/EGFRvIII-Targeted HSV to Key Neutralizing Antibodies
- PMID: 32042851
- PMCID: PMC7000558
- DOI: 10.1016/j.omtm.2019.12.013
Point Mutations in Retargeted gD Eliminate the Sensitivity of EGFR/EGFRvIII-Targeted HSV to Key Neutralizing Antibodies
Abstract
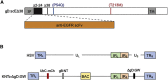
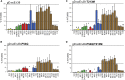
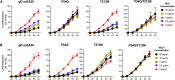
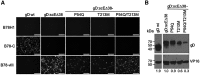
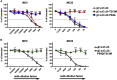
Effective oncolytic virotherapy may require systemic delivery, tumor targeting, and resistance to virus-neutralizing (VN) antibodies. Since herpes simplex virus (HSV) glycoprotein D (gD) is the viral attachment/entry protein and predominant VN target, we examined the impact of gD retargeting alone and in combination with alterations in dominant VN epitopes on virus susceptibility to VN antibodies. We compared the binding of a panel of anti-gD monoclonal antibodies (mAbs) that mimic antibody specificities in human HSV-immune sera to the purified ectodomains of wild-type and retargeted gD, revealing the retention of two prominent epitopes. Substitution of a key residue in each epitope, separately and together, revealed that both substitutions (1) blocked retargeted gD recognition by mAbs to the respective epitopes, and, in combination, caused a global reduction in mAb binding; (2) protected against fusion inhibition by VN mAbs reactive with each epitope in virus-free cell-cell fusion assays; and (3) increased the resistance of retargeted HSV-1 to these VN mAbs. Although the combined modifications of retargeted gD allowed bona fide retargeting, incorporation into virions was partially compromised. Our results indicate that stacking of epitope mutations can additively block retargeted gD recognition by VN antibodies but also that improvements in gD incorporation into virus particles may be required.
Keywords: glycoprotein D; neutralizing antibodies; oncolytic HSV; systemic treatment; tumor targeting.
© 2020 The Authors.
Figures
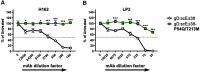
Similar articles
-
Syncytial Mutations Do Not Impair the Specificity of Entry and Spread of a Glycoprotein D Receptor-Retargeted Herpes Simplex Virus.J Virol. 2016 Nov 28;90(24):11096-11105. doi: 10.1128/JVI.01456-16. Print 2016 Dec 15. J Virol. 2016. PMID: 27707922 Free PMC article.
-
Antibody Screening System Using a Herpes Simplex Virus (HSV)-Based Probe To Identify a Novel Target for Receptor-Retargeted Oncolytic HSVs.J Virol. 2021 Apr 12;95(9):e01766-20. doi: 10.1128/JVI.01766-20. Print 2021 Apr 12. J Virol. 2021. PMID: 33627393 Free PMC article.
-
Dual Ligand Insertion in gB and gD of Oncolytic Herpes Simplex Viruses for Retargeting to a Producer Vero Cell Line and to Cancer Cells.J Virol. 2018 Feb 26;92(6):e02122-17. doi: 10.1128/JVI.02122-17. Print 2018 Mar 15. J Virol. 2018. PMID: 29263257 Free PMC article.
-
Oncolytic Herpes Simplex Virus Vectors Fully Retargeted to Tumor- Associated Antigens.Curr Cancer Drug Targets. 2018;18(2):162-170. doi: 10.2174/1568009617666170206105855. Curr Cancer Drug Targets. 2018. PMID: 28176649 Review.
-
Retargeting Strategies for Oncolytic Herpes Simplex Viruses.Viruses. 2016 Feb 26;8(3):63. doi: 10.3390/v8030063. Viruses. 2016. PMID: 26927159 Free PMC article. Review.
Cited by
-
Novel mutations in UL24 and gH rescue efficient infection of an HSV vector retargeted to TrkA.Mol Ther Methods Clin Dev. 2023 Jul 3;30:208-220. doi: 10.1016/j.omtm.2023.06.012. eCollection 2023 Sep 14. Mol Ther Methods Clin Dev. 2023. PMID: 37519407 Free PMC article.
-
Oncolytic herpes simplex viruses designed for targeted treatment of EGFR-bearing tumors.Mol Ther Oncol. 2024 Jan 10;32(1):200761. doi: 10.1016/j.omton.2024.200761. eCollection 2024 Mar 21. Mol Ther Oncol. 2024. PMID: 38596286 Free PMC article.
-
Modelling the spatial dynamics of oncolytic virotherapy in the presence of virus-resistant tumour cells.PLoS Comput Biol. 2022 Dec 6;18(12):e1010076. doi: 10.1371/journal.pcbi.1010076. eCollection 2022 Dec. PLoS Comput Biol. 2022. PMID: 36473017 Free PMC article.
-
Current strategies to circumvent the antiviral immunity to optimize cancer virotherapy.J Immunother Cancer. 2021 Apr;9(4):e002086. doi: 10.1136/jitc-2020-002086. J Immunother Cancer. 2021. PMID: 33795384 Free PMC article. Review.
-
Virus Against Cancer: Paradigm-Shifting Biological Concepts.Curr Neurol Neurosci Rep. 2025 Jun 14;25(1):41. doi: 10.1007/s11910-025-01428-4. Curr Neurol Neurosci Rep. 2025. PMID: 40515944 Review.
References
-
- Andtbacka R.H., Kaufman H.L., Collichio F., Amatruda T., Senzer N., Chesney J., Delman K.A., Spitler L.E., Puzanov I., Agarwala S.S. Talimogene laherparepvec improves durable response rate in patients with advanced melanoma. J. Clin. Oncol. 2015;33:2780–2788. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

